-
AI based chat application for life sciences:
Part II role of ontologies

Explore the vital role of ontologies in enhancing AI-based chat applications for life sciences, with a focus on improving transparency and data meaning.
Read -
Ontology mapping:
Finding the right automated approach

Ontologies are crucial for unlocking information. However, similar types have been created for different needs, reducing their interoperability. In this blog, we look at some of the automated approaches for large-volume ontology mapping.
Read -
Novelty in life science: Looking into the unseen


In life science research, navigating the complexities of innovation is crucial for breakthroughs. SciBite’s Novelty model, a sophisticated Machine Learning classifier, distinguishes true innovation in scientific texts.
Read -
AI based chat application for life sciences:
Part I key considerations

Are your teams now posing potentially confidential questions to consumer tools such as Bard and ChatGPT, relying on their responses?
Read -
Ontology mapping:
Advancing data interoperability

Ontologies are crucial in unlocking information that helps fuel innovation. However, similar ontologies have emerged within the same domain, making it difficult for researchers to identify the right ontology to choose. Here we look at some of the complexities and strategies for handling ontology mappings.
Read -
FOSHU: The original functional foods


Explore how FOSHU informed the creation of our latest VOCab, a valuable asset for research in food science and dietary therapeutics. We begin by taking a deeper dive into Japan’s diverse FOSHU market
Read -
With new tech comes new operationalization considerations


As technology advances, the landscape of operationalization undergoes a profound shift. Here, we unravel the intricacies that accompany new tech, exploring key operationalization considerations shaping the realms of machine learning and semantic indexing.
Read -
Functional foods: Revolutionizing health through diet and innovation


Embark on a journey through the transformative realm of functional foods, where the convergence of nature's wisdom and cutting-edge innovation is reshaping our approach to health. In the 21st century, these foods emerge as powerful allies, combating lifestyle diseases and ushering in a new era of well-being.
Read -
What is Retrieval Augmented Generation and why is the data you feed it so important?


Within the life sciences, evidence-based decision-making is imperative; wrong decisions can have dire consequences. As such, it is vital that systems that support the generation and validation of hypotheses provide direct links, or provenance, to the data that was used to generate them. But how can one implement such a workflow?
Read -
Unlocking important RWE from patient data (Part 3) – Can we find all the relevant patients?


In our final installment of this series, we demonstrate how to extract a relevant subset of patients from the simulated data using two approaches – one using SciBite tools and one without.
Read -
FAIR data – Ten simple rules to FAIRify your data


In the fourth and final blog in this series Scibite’s Head of Ontologies, Jane Lomax, shares her top 10 simple rules to start and progress your FAIR data journey.
Read -
Are ontologies still relevant in the age of LLMs?


Technological advancements exhibit varying degrees of longevity. Some are tried and trusted, enduring longer than others, while other technologies succumb to fleeting hype without attaining substantive fruition. One constant, in this dynamic landscape is the data.
Read -
Microbiome repurposing: Is it a potential therapeutic approach & how can we do it with LLMs?


Discover the past and future of microbiome-based healing. From ancient remedies to modern AI, learn how SciBite's groundbreaking approach blends Large Language Models (LLMs) with advanced tech to unravel the potential of therapeutic microbiomes.
Read -
Girls in Tech – SciBite/Elsevier initiative to inspire the next generation of young women in tech


In a world where technology plays an increasingly pivotal role in shaping our lives, it is crucial to recognize the contributions of women and the importance of empowering the next generation of female tech professionals.
Read -
FAIR as a means to get more value from your data


In this blog, we’ll explore a selection of the many ways organizations can leverage the rapid developments in data discovery, machine learning, and data mining to release value from this asset.
Read -
Unlocking important real world evidence from patient data (Part 2) – Data domain deep dive


In this part of our blog series, "Unlocking important real-world evidence from patient data," we will demonstrate our expertise in various important data domains using SciBite tooling, including problem list diagnoses, lab orders, and medication orders.
Read -
The key to being FAIR


In our previous blog, we explained why FAIR data is important not only for biotech and pharmaceutical companies but also for their partners. Here we describe how ontologies are the key to having the richly described metadata that is at the heart of making data FAIR. Let’s explore how ontologies help with each aspect of the FAIR data principles…
Read -
Unlocking important RWE from patient data (Part 1) – Why and how?


In this three-part blog series, we explore the challenges healthcare organizations face in unlocking patient data for real-world evidence. In part 1 Unlocking Important Real World Evidence (RWE) from Patient Data – Why and How?
Read -
Why do you need FAIR data?


For many organizations, the idea of adopting FAIR can be confusing and daunting. Over the coming weeks, we’ll present a series of blogs to help demystify FAIR. In this series, we’ll cover topics including how ontologies provide the key to being FAIR, and how FAIR enables you to get more value from your data.
Read -
How SciBite and Elsevier manage KOL identification


Identifying KOLs enables our customers to be the first to follow the latest trends and markets or start new collaborations. As you can imagine, spotting and engaging KOLs as fast and accurately as possible is crucial - read more to understand how.
Read -
Large language models (LLMs) and search; it’s a FAIR game


Large language models (LLMs) have limitations when applied to search due to their inability to distinguish between fact and fiction, potential privacy concerns, and provenance issues. LLMs can, however, support search when used in conjunction with FAIR data and could even support the democratisation of data, if used correctly…
Read -
A report from the Biocuration 2023 Conference in Padua, Italy


The Biocuration Conference this year was held in the beautiful historic town of Padua in the Veneto region of Italy, renowned for its ancient University and picturesque old town. The stylish and relaxed atmosphere was the perfect place for catching up with old friends and establishing new connections and collaborations.
Read -
Double recognition for SciBite in Bio-IT World Innovative Practices Awards 2023


SciBite has won the Innovative Practice Award at Bio-IT World 2023 for its collaboration with the City of Hope and also for its pivotal role as the ontology ecosystem in AbbVie’s Innovative Practices Award win.
Read -
A review of the Pistoia Alliance Spring Conference 2023


Last week SciBite was lucky enough to attend, and present at, the Pistoia Alliance Annual Spring Conference ‘23, held at the fantastic Leonardo Royal Hotel, St. Pauls, London. Read the thoughts of our Director of Technical Consultants, Joe Mullen
Read -
Toxicology keeps us safe using scientific evidence


What do medical devices and medicines, food products and cosmetics, household products and even smoking devices have in common? They all contain multiple ingredients, from natural to synthetic ones, in different quantities, all of which will be exposed to the population.
Read -
Training AI is hard, so we trained an AI to do it


GPT3 is a large language model that is capable of generating text with very high fidelity. Unlike previous models, it doesn't stumble over its grammar or write like an inebriated caveman. In many circumstances it can easily be taken for a human author, and GPT-generated text is increasingly prolific across the internet for this reason.
Read -
Streamlining data-intensive scientific workflows through FAIR data


Streamlining data-intensive scientific workflows and supply chains through FAIR data, data models and applications – A collaboration between L7 Informatics and SciBite. With increasingly complex manufacturing and supply chains in the life sciences, there is a requirement for flexible and extensible tools to support data management.
Read -
Revolutionizing Life Sciences: The future of AI in Life Science [Part 2]


As discussed in part 1, Artificial intelligence (AI) has revolutionized several areas in life sciences, including disease diagnosis and drug discovery. In this second blog, we introduce some specific text-based models whilst also discussing the challenges and future impact of AI in Life Science.
Read -
SciBite launches Workbench –
Taking the effort out of tabular data curation

SciBite, a leading provider of semantic technology solutions, has today announced the launch of Workbench, a structured data annotation tool that simplifies the process of curating data to terminology and ontology standards.
Read -
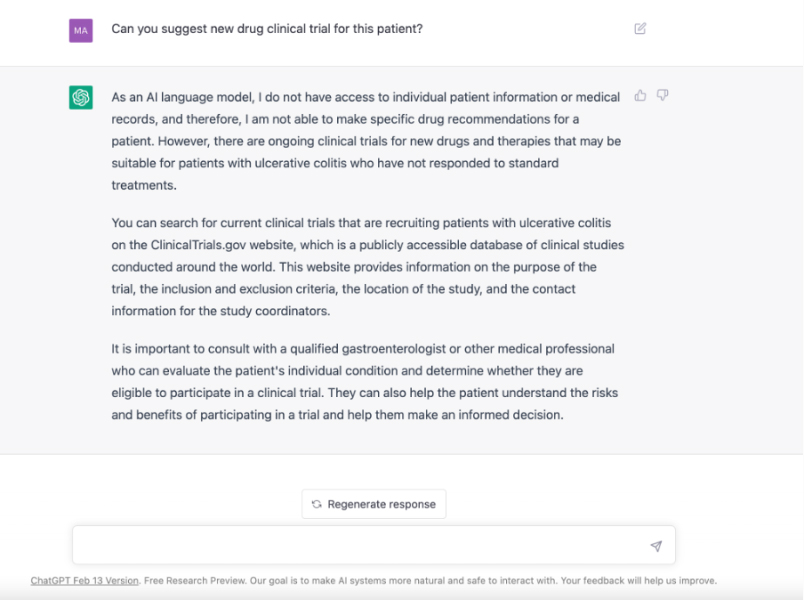
Matching patients to clinical trials


Patient X, suffering from an untreatable gastrointestinal disease, chats with a large language model for advice. GPT suggests looking at clinical trials and Patient X finds 10 active recruiting trials but is unsure which to choose. Patient X consults his doctor, who recommends a trial from a pharmaceutical company. What could go wrong?
Read -
Revolutionizing Life Sciences: The incredible impact of AI in Life Science [Part 1]


Artificial intelligence (AI) has been applied to numerous aspects of the life sciences, from disease diagnosis to drug discovery; in the first of this two-part blog series, we outline the impact of AI in Life Science and illustrate the various success stories of AI in Life Science.
Read -
IDBS and SciBite bring enterprise ontologies and scientific data management together


IDBS, a leading provider of life science informatics and process data management solutions, and SciBite, an award-winning provider of semantic analytics technologies, today announce a new partnership to shift the way research data platforms approach FAIR (Findable, Accessible, Interoperable, and Reusable) data.
Read -
Healthcare digital transformation challenges: Can we enable healthcare systems to trust their data?


At SciBite, we are passionate about enabling organizations to make full use of their data to help them make evidence-based decisions, especially to help organizations overcome their healthcare digital transformation challenges. To support organizations on this journey, we offer a suite of products to help organizations adopt FAIR data standards.
Read -
SciBite unveils partnership with Modak


Modak, a leading provider of data engineering solutions, and SciBite, an award-winning provider of semantic analytics technologies, today announced a partnership that will empower Life Science enterprises to fast track the process of generating insights from research publications, patents, and documents, which is crucial for advancing scientific discovery.
Read -
How SciBite technology can facilitate gene-disease relationship extraction


As genomic sequencing technologies get more advanced, large numbers of gene-disease associations have emerged. A gene with an unclear role within a disease is a source of ambiguity and can lead to misdiagnosis. In this blog, we demonstrate how semantic search technology can facilitate Gene-Disease Relationship Extraction.
Read -
Harnessing our latest VOCab:
Emtree

The 6.5.2 release of SciBite’s VOCabs introduces a range of new VOCab packs as well as updates to existing vocabularies. In this blog series we’ll be introducing each of the new VOCabs: IDMP, a new Sequence Ontology VOCab as part of the Genotype-Phenotype VOCab pack and, first up, the new Emtree VOCab pack.
Read -
What is IDMP – Navigating the ISO IDMP standards with SciBite ontologies


As of the first quarter of 2023, it will be mandatory for pharmaceutical manufacturers that market to the EU to comply with the European Medicines Agency (EMA) ISO Standards for the Identification of Medicinal Products (IDMP).
Read -
Creating a SciBite VOCab from a public ontology


Public ontologies are essential for applying FAIR principles to data but are not built for use in named entity recognition pipelines. At SciBite, we build on the public ontologies to create VOCabs optimized for NER. In this blog, discover how we create a SciBite VOCab from a Public Ontology.
Read -
Why use your ontology management platform as a central ontology server


Raw data has the inherent characteristic of being unstructured with potential quality issues such as inaccurate, incomplete, inconsistent, and duplicated. Therefore, it must be processed before it can be used for subsequent analysis and confident data-driven decisions. This is where ontologies come into play.
Read -
Leveraging semantics for effective navigation of scientific content


A Copyright Clearance Center & SciBite Perspective - In an increasingly data-driven society, it can be overwhelming to keep your knowledge base current and effectively utilize data. The result in many organisations is underutilising the data and content available to them.
Read -
What’s in our 6.5.2 TERMite / VOCabs release


SciBite’s vocabularies fuel a host of use cases, from complex querying to data integration and discovery of new knowledge. In the 6.5.2 release of VOCabs, SciBite introduces the new Emtree VOCab pack, as well as a new Sequence Ontology vocab to the Genotype-Phenotype vocab pack. Several updates to existing vocabularies are also included.
Read -
Delivery of precision medicine through alignment of clinical data to ontologies


Precision medicine is changing the way that we think about the treatment of disease, moving from broad-acting therapies to therapies tailored to the individual patient. This increasingly relies on real-world data (RWD), encompassing a diverse range of sources, spanning multi-omic molecular characterisation of the patient’s condition, clinical presentation, treatment, and broader medical histories.
Read -
SciBite named ‘Best of Show’ at Bio-IT World 2022 for CENtree


SciBite’s ontology management platform CENtree won ‘Best of Show” at this year’s Bio-IT World 2022. Build the foundations for data sharing with our collaborative ontology management platform.
Read -
SciBite’s newest VOCabs – What’s in our 6.5 release


SciBite’s VOCabs power a host of semantic use cases including search and analytics. Created from public ontologies and reference databases for a wide range of topics, these vocabularies are enriched using our proprietary tools, and curated by our team of experts to ensure maximum capture of relevant synonyms by subject area and context.
Read -
CODiE 2022 SIIA finalist in best Integration Platform as a Service (iPaaS)


SciBite's ontology management platform, CENtree earns nod from industry leaders by being named a 2022 SIIA CODiE Award finalist in the Best Integration Platform as a Service category.
Read -
SKOS in CENtree: Further support in our latest 2.1 release


At SciBite terminologies underpin all that we do. There are many ways to represent and build a standardised terminology, each with different levels of complexity. On one hand you have simple, informal, lightweight terminologies (e.g., glossaries, dictionaries, and thesauri), where the meaning (semantics) of terms is captured using natural language.
Read -
SciBite brings enterprise ontologies to Benchling – Ontology backed data capture


Unstructured and siloed data in the life sciences remains a significant barrier to fulfilling the promise of digital transformation. Awareness is growing for the importance of data capture and storage, enabling it to be effectively found, accessed, used interoperably and reused. These are the foundations of FAIR. Capturing data with FAIR in mind, ensuring your data is “born FAIR”, is key to unlocking the full potential of data.
Read -
SciBite announces the release of SciBite Search 2.0


In this blog we announce the v2.0 release of SciBite Search, our intelligent scientific search platform. We’ve expanded our Elsevier data connectivity, broadening the sources you can load and search, as well as a host of features that improve the user experience.
Read -
SciBite announces the release of CENtree 2.0.1


In this blog we announce the 2.0.1 release of CENtree, SciBite’s ontology management platform, which sees the introduction of features that enable you and your team greater control over managing and deploying ontologies in your applications, and a closer integration with TERMite.
Read -
SciBite and Sinequa join forces to transform scientific search


SciBite and Sinequa's new collaboration combines custom ontologies with a powerful search platform to help researchers find answers fast.
Read -
SciBite and Stardog build a Knowledge Graph for drug discovery


In a recent webinar, SciBite and Stardog team members demonstrated how to build a knowledge graph to identify candidate drugs for a rare disease.
Read -
Introducing efficiencies into the scientific reading experience


First and foremost, researchers need access to a corpus of scientific literature and secondly, they need a robust and reproducible way to search that content. Through partnership, SciBite and Copyright Clearance Center bring together scientific content and advanced search capabilities.
Read -
SciBite launches SaaS versions of its semantic technology products


SciBite has today unveiled its new Software-as-a-Service (SaaS) version of TERMite, SciBite’s named entity recognition engine, and its CENtree™ ontology management system.
Read -
SciBite and L7 Informatics partner to deliver data ontology search through L7|ESP™


SciBite and L7 Informatics announce joint partnership to support discovery and collaboration in the life sciences.
Read -
The powerful combination of semantics-based Machine Learning and domain expertise


In this blog hear about SciBite's recent talk at Pistoia Alliance’s Spring Virtual Conference on semantics-based machine learning and domain expertise on a day dedicated to emerging science and technologies.
Read -
SciBite Search: Next-generation scientific search & analytics platform


SciBite launches AI-driven semantic search platform to help manage data deluge. Intelligent scientific search platform, SciBite Search, enables researchers to quickly find meaningful insights from structured and unstructured public and proprietary biomedical data.
Read -
Addressing common challenges with Knowledge Graphs


In this blog we describe the pivotal role of semantic enrichment in the creation of effective Knowledge Graphs, and illustrate how semantic Knowledge Graphs help answer complex scientific questions.
Read -
Exploring breast cancer biomarkers with a literature biomarker database
In this piece we'll show how natural language processing can be applied to build a searchable database of disease biomarkers, presented in the context of their corresponding scientific publications. To illustrate the power of this approach we'll focus on examples of protein biomarkers relating to Breast Cancer.
Read -
Using the SciBite knowledge graph to explore biomedical literature
“Do you have a pre-made knowledge graph covering biomedical literature?” is a question we often hear at SciBite. The answer is yes we do, and in this blog post we’ll describe what our SciBite Knowledge Graph is, its content and the types of questions it can answer.
Read -
Annotation of the Covid-19 open research dataset for the scientific research community
In this blog find out how the SciBite team has responded to the tech community call to arms from The White House after they released an Open Research Dataset (CORD-19), with the hope to help uncover insights and answer high-priority scientific questions related to Covid-19.
Read -
What’s new in CENtree 1.4:
Making ontology management simpleFind out what's new in CENtree 1.4, the latest release of the enterprise ready multi-user ontology management platform for browsing and managing ontologies.
Read -
Bringing FAIR data and CMC procedures together
In this blog we introduce our new package of vocabularies designed to enable the FAIR data principles and help pharmaceutical companies navigate their documents with respect to Chemistry, Manufacture and Control (CMC) procedures.
Read -
Elsevier acquires SciBite to accelerate solutions for life sciences and corporate R&D industries
Elsevier, a global research publishing and information analytics provider, and part of RELX, has acquired SciBite, a semantic AI company headquartered in Cambridge, UK, to help customers make faster, more effective R&D decisions through advanced text and data intelligence solutions.
Read -
Sprinkling a little semantic enrichment into your data catalog
This blog focuses on the use and value of data catalogs and Master Data Management (MDM) tools and how the additional layer of Semantics is required in order to truly see their value for enterprises looking to manage their data better.
Read -
Unlocking patents as a data source in the Life Sciences
Throughout this blog we highlight some complexities that exist in extracting meaningful information from patents and show various solutions, making use of SciBite technology alone or, augmented by or delivered by our partners.
Read -
SciBite wins Queen’s Award: Enterprise in Innovation
SciBite, the data first, semantic analytics software company, today announced being named as a winner of the Queen’s Award for Enterprise in Innovation – the highest official UK awards for businesses.
Read -
Using scientific data technology to access the emerging trends in immuno-oncology research
In this blog, we explain more about our Immunonc vocabulary, comprising the concepts relating to this field, which enables our customers to be better informed about the current advances and emerging trends in cancer immunotherapy research, keeping them at the forefront of cancer treatment.
Read -
SciBite announced as fastest-growing company in Pharma Fast 50
SciBite was named the fastest-growing company identified in this year’s Alantra Pharma Fast 50, an annual ranking of privately-owned pharmaceutical businesses in the UK based on growth speed.
Read -
The allotrope ontologies: The power of open ontologies in industry
In this blog, find out how our next-generation ontology management tool CENtree can be used to manage the allotrope ontologies.
Read -
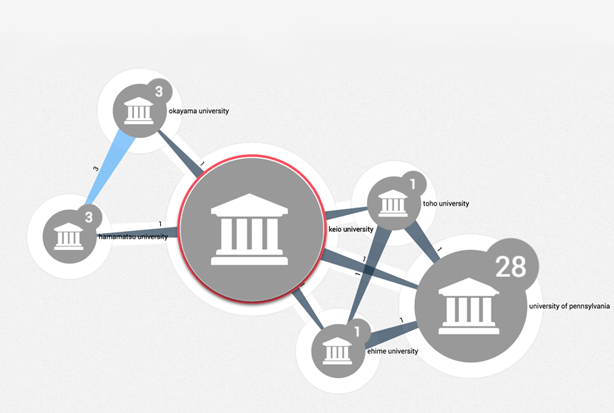
Machine Learning insights from Japanese language academic text
In this blog, we delve into how we applied novel machine learning and curation methods to Japanese language literature, techniques we believe are transferable to other under-supported languages.
Read -
What is a Semantic Knowledge Graph?
At a time where more and more of our customer projects revolve around knowledge graph creation, we thought it was about time we blogged on what exactly a knowledge graph is and explain a bit more about how our semantic enrichment technology is being used to facilitate the production of such a powerful data model.
Read -
Semantic approach to training ML data sets using ontologies & not Sherlock Holmes
In this blog we discusses how Sherlock Holmes (amongst others) made an appearance when we looked to exploit the efforts of Wikipedia to identify articles relevant to the life science domain for a language model project.
Read -
Building the future of text analytics
SciBite CSO and Founder Lee Harland features in KM World Magazine, where he talks about the future of text analytics and how ontologies are the de facto standard to encode semantics in an understandable form for both humans and machines.
Read -
Using Enterprise Search to unlock the wealth of R&D data
Learn more in this blog post about our partnership with Sinequa and how our technologies work together to provide a winning combination of Cognitive Search and powerful Life Sciences Semantics, as we explain how enterprise search platforms are enabling Pharmaceutical companies to become more information-driven.
Read -
A helping hand from BERT for Deep Learning approaches
SciBite CSO and Founder Lee Harland shares his views on the use of BERT (or more specifically BioBERT) for deep learning approaches.
Read -
Powering question-driven problem solving with semantic integration
In this blog post hear from GSK's Scientific Lead within the Data and Computational Sciences Solutions team, Samiul Hasan on how semantic integration can be made to ultimately become part of an integrated learning framework for more informed scientific decision making.
Read -
The benefits of semantically enriching document mining for chemists
In this blog post, learn more about how our partner ChemAxon have integrated SciBite’s ultrafast named entity recognition (NER) and extraction engine solution, TERMite, into their leading cheminformatics platform, and how this can benefit your organisations informatics architecture.
Read -
Using ontologies to unlock the full potential of your scientific data – Part 2
This blog post focuses on mapping, building, and managing ontologies. In my previous blog, I described what ontologies are and how you can use them to make the best use of scientific data within your organization. Here I’ll expand upon this and focus on mapping, building, and managing ontologies.
Read -
Using ontologies to unlock the full potential of your scientific data – Part 1
In the first of this two-part blog, I describe what ontologies are and how you can use them to make the best use of scientific data within your organisation.
Read -
The pivotal role of semantic enrichment in the evolution of data commons
In this blog post, discover how Pfizer have integrated SciBite’s semantically enriched vocabularies into their Data Commons project, which has the goal of enabling scientists to develop and refine hypotheses by investigating correlations between genetic and phenotypic data.
Read -
How ontology enrichment is essential in maintaining clean data
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the final blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read -
The importance of facilitating collaboration and integration
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the third blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read -
Why simplifying visualization and curation is better for everyone
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the second blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read -
The benefits of centralizing ontology management
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the first blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read -
How the use of Machine Learning can augment adverse event detection
When it comes to identifying adverse events (AEs), things are not always as they seem. Consider a paper describing a new treatment for a given illness - how can we determine which adverse event terms refer to actual adverse events as opposed to symptoms of the illness itself, given that those terms may be identical? Is this new drug treating arrhythmias or causing them, for example?
Read -
SciBite wins Queen’s Award: Enterprise in International Trade
SciBite today announced being named as a winner of the Queen’s Award: Enterprise in International Trade – the highest official UK awards for businesses.
Read -
SciBite launches CENtree, ontology management for life sciences
Cambridge, UK - SciBite, the award-winning semantic technology company, today announced the launch of CENtree, an innovative, collaborative platform which revolutionizes the way life sciences organizations manage and release ontologies.
Read -
SciBite announced as best of show award finalists for Bio-IT World 2019
SciBite has been shortlisted for Bio-IT World 2019’s prestigious Best of Show Award.
Read -
SciBite Ontology Services offers expert ontology solutions
SciBite, the award-winning semantic technology company, today announced the launch of SciBite Ontology Services.
Read -
SciBite announced as finalists in Bio-IT World 2019’s Innovative Practices Awards
SciBite's innovative project with LifeArc has been shortlisted for Bio-IT World’s inaugural Innovative Practices Awards 2019.
Read -
How biomedical ontologies are unlocking the full potential of biomedical data
Our latest blog explains how SciBite's Ontologies team takes public biomedical ontologies and tailors them so that they can be used for named entity recognition (NER).
Read -
How semantic enrichment technology is changing the way we search
In our latest blog we discuss the challenges life sciences companies, like LifeArc, face in keeping up-to-date with scientific literature, and how semantic enrichment technology can automate this process to reduce the time spent mining data by up to 80%.
Read -
SciBite joins Accenture Life Sciences Ecosystem
SciBite joins Accenture’s open partner ecosystem, designed to help independent software vendors (ISVs) and life science companies team more effectively to accelerate drug discovery efforts and improve patient outcomes.
Read -
SciBite & RDF (Resource Description Framework) – A natural semantic fit
In this article, we’ll explore how SciBite’s platform works with the semantic web and its primary data representation format, RDF, along with the benefits each technology brings.
Read -
Are ontologies relevant in a machine learning-centric world?
SciBite CSO and Founder Lee Harland shares his views on why ontologies are relevant in a machine learning-centric world and are essential to help "clean up" scientific data in the Life Sciences industry.
Read -
Text mining drug labels for genetic factors influencing efficacy and safety
Only 50-75% of patients respond beneficially to first-line drug therapy, causing unnecessary costs to healthcare providers and, more importantly, adversely impacting a patient’s quality of life. Text mining drug labels for genetic factors influencing efficacy and safety to support clinicians at the point of prescription.
Read -
SciBite and Hadoop: Transforming Big Data
In this post we explore how SciBite's semantic platform fits into these infrastructures and how it can be used to unlock the knowledge held in text form within an organisation to power next-generation analytics and insight.
Read -
Ontology mapping: You say tomato I say Solanum Lycopersicum…
In this post, we explore this important but often overlooked topic and discuss its relevance to the work of SciBite and other groups, such as FAIR and the Pistoia Alliance within the broader scientific community.
Read -
Of burns and bums: Machine Learning surprises!
As many of our regular visitors will know, the focus of our work here at SciBite is unlocking the knowledge held in the vast amount of biomedical text researchers have access to. Sometimes this yields well, interesting, results...
Read -
SciBite and ONTOFORCE partner to make semantic search even more powerful
Cambridge, UK – SciBite, a firm whose technology is disrupting the traditional approaches of working with scientific information across the globe, today announced a strategic partnership with ONTOFORCE, the leading data integration company.
Read -
SciBite and ChemAxon to deliver an integrated solution for biology and chemistry research
SciBite, announces strategic partnership with ChemAxon, the leading cheminformatics company, enabling Pharmaceutical companies to capitalise on unlocking the wealth of R&D data that is at their disposal.
Read -
Drug repurposing, rare diseases and semantic analytics
In this blog we cover how to look potentially reduce the cost of and speed up the repurposing pipeline.
Read -
A hacker’s guide to understanding bio-ontology jargon
Perfect for those new to bio-ontologies or who work with ontologists - a whole new vocabulary deciphered!
Read -
CCC launches RightFind Insight, transforming scientific research
Just released by the Copyright Clearance Center, a semantic search solution applied to full-text articles
Read -
Taking semantic search to full text
Guest post from CCC, leader in creating global licensing and full content solutions, on the advantage of full text for semantic search.
Read -
GSK Japan selects semantic platform to enhance pharmacovigilance capabilities
SciBite today announced that GSK Japan, one of Japan’s leading research-based pharmaceutical and healthcare companies, has selected SciBite’s Semantic Platform to enhance pharmacovigilance capabilities and deliver on its commitment to improve the quality of human life.
Read -
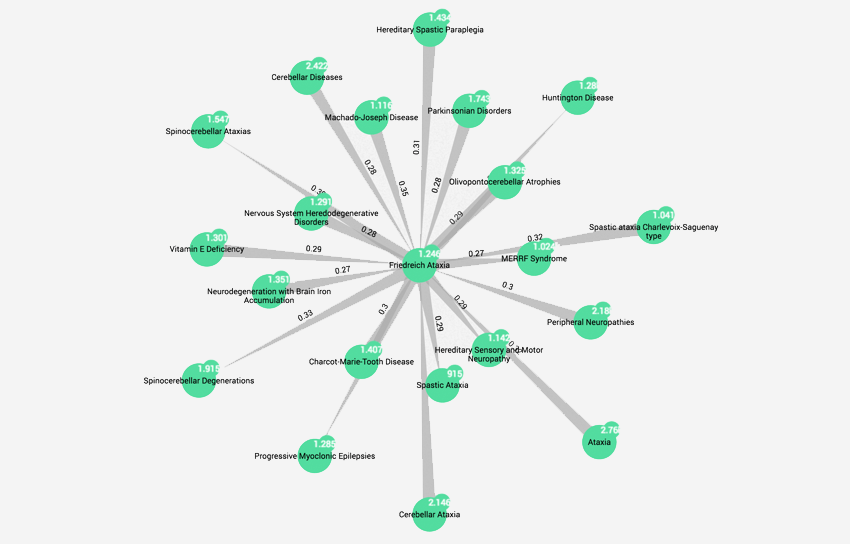
Graph technologies to help solve the challenges of rare diseases research
Rare Disease Day took place on 28 February, and to help publicise it, we thought we’d raise awareness of the kind of work happening in this area within the pharmaceutical industry, and how big data tools are impacting those efforts.
Read -
Exploring mechanistically-related diseases through shared phenotypic profiles
Disease detective part 2: Today, we’ll look at a fresh way of enabling scientific researchers, either in pharmaceutical R&D or in medical institutes to deepen their investigations and consider new links.
Read -
Rare disease collaboration networks
Disease Detective Part 1: In celebration of Rare Disease Day 28th Feb, we have a 3 part blog post looking into some of the challenges/analysis techniques involved in the research process.
Read -
SciBite and PerkinElmer provide advanced analytics from unstructured scientific data
PerkinElmer, Inc., today announced sophisticated scientific semantic enhancements to the PerkinElmer Signals™ Perspectives platform, powered by SciBite and Attivio®.
Read -
SciBite partners with Copyright Clearance Center
We live in a world of increasing volumes, sources and formats of scientific content. So how can you find what is important quickly, simply, and with the confidence that you have missed nothing?
Read -
SciBite becomes new Pistoia Alliance member
SciBite are delighted to join the Pistoia Alliance helping to address some of the challenges faced with big data in Global Life Sciences R&D. We look forward to meeting all the attending members at the European Conference starting tomorrow.
Read -
The Relationship Game – Knowledge Graphs
Scientific knowledge can be represented as relationships between things. Thousands or millions of such relationships make a knowledge graph or network analysis. SciBite technology enables extraction of these relationships, and in doing so, can uncover knowledge that might otherwise have remained hidden
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.