
Why use your ontology management platform as a central ontology server
 |
 |
Raw data has the inherent characteristic of being unstructured with potential quality issues such as inaccurate, incomplete, inconsistent, and duplicated. Therefore, it must be processed before it can be used for subsequent analysis and confident data-driven decisions. This is where ontologies come into play.

Just a quick recap first. Raw data has the inherent characteristic to be unstructured with potential quality issues such as inaccurate, incomplete, inconsistent, and duplicated. Therefore, it must be processed before it can be used for subsequent analysis and confident data-driven decisions.
This is where ontologies come into play.
So why do we need ontologies?
Ontologies provide a way to codify or standardize the business language of organizations across a domain in a machine-readable form. They enable the information sharing between diverse systems within the same domain. In the life sciences domain, some of the terms (entities) of interest include molecules, targets, diseases, tissues, cell lines, anatomical terms, species, strains to name a few.
Those entities may have associated properties (such as labels, definition, synonyms, mappings) and relationships (such as ‘part of’, ‘derives from’, ‘develops from’) to another entities or entity’s categories. Thus, a drug molecule can interact with a target and modulate its activity (a relationship between different entities) or may be an analgesic, a corticosteroid, an expectorant, a sedative… (a category).
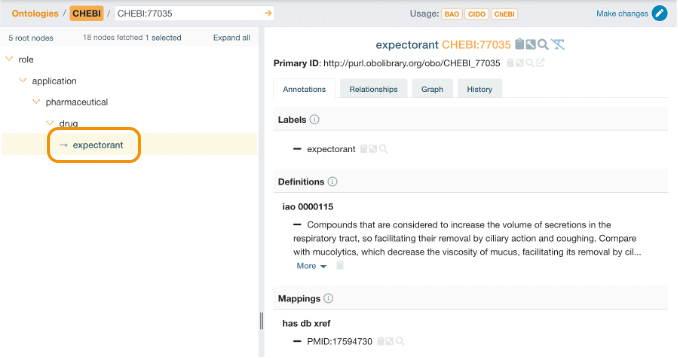
Figure 1: Example of drug’s category from the Chemical Entities of Biological Interest (ChEBI) ontology.
Each entity has both a unique identifier as well as potentially multiple externally issued ones (such as drug INN, CAS, SMILES, InChI) depending on the queried ontology.

Figure 2: Examples of externally issued drug’s identifiers from the ChEBI ontology.
The categories can be thought of as adjectives, while the relationships can be thought of as prepositional phrases (such as ‘is enantiomer of’, ‘is conjugate acid of’).
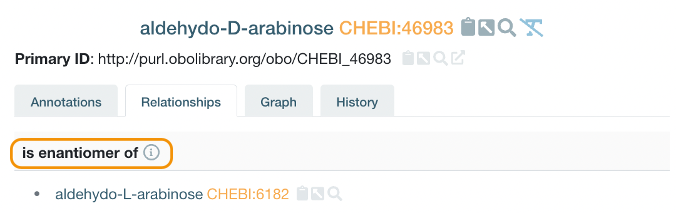
Figure 3: Example of chemical entity’s relationship from the ChEBI ontology.
Using an ontology server for ontology development
An ontology defines a set of classes, attributes (or properties) and relationships with which to model a domain of knowledge (please read this article for understanding ontology jargon). With the increased adoption of ontologies in the life sciences domain, CENtree plays a fundamental role in data standardization and interoperability.
Standardization is the result of using ontologies to eliminate ambiguity from technical languages and thus enabling communication and knowledge sharing between agents either human or software thanks to resolvable globally unique and persistent identifiers. It also increases compatibility and interoperability across data sets allowing information to be shared within a larger network (e.g., collaboration, partnerships).
CENtree aims to facilitate ontology development, edition and visualization but it can also be used as an ontology’s server. It provides an intuitive Web interface for displaying the details and hierarchy of a specific ontology term.
Maintaining ontologies in an ontology server
There are a couple of Web sites which provide lists of scientific ontologies (e.g., OBO Foundry, BioPortal). When an application (e.g., an ELN, a data catalog, a knowledge graph) needs to consume some of the ontologies, it is required to learn each individual underlying API and manage the authentication mechanism for each of them.
With CENtree, this process is simplified and quite straightforward. Like all applications developed at SciBite, CENtree is an API-first application. This means that we developed exhaustive collections of API endpoints first, and then, we designed the Web user interface (UI) based on those collections. This also means that all the actions, doable in the UI, can be done using the API endpoints (here, some examples from the “search-resource” and “ontology-metadata-resource” collections).

So, to integrate the ontologies that are hosted and maintained in CENtree, a unique API knowledge is required as well as a single authentication! This represents a huge benefit over the multitude of individual APIs of the different public ontologies. The ROI of leveraging CENtree as an ontology server is quite immediate in terms of efforts and resources required to integrate the required ontologies in a downstream application.
The UI of CENtree has been designed to be accessible to a broad audience and not just experts (e.g., ontologists). Loading public ontologies is a one-click process for CENtree users having the appropriate permissions, the related API endpoint being:

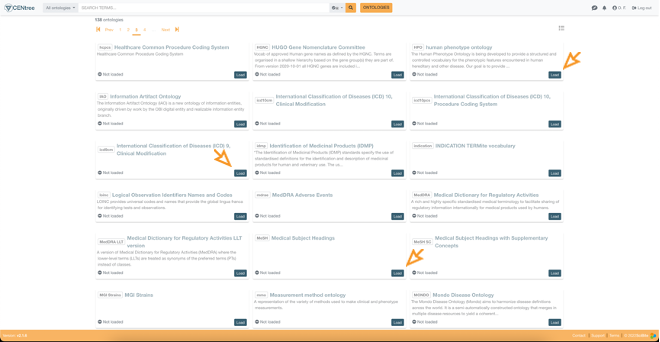
Figure 4: One-click feature to load public ontologies in the CENtree UI.
Benefits of using CENtree as an ontology server
Here are some of the benefits of using CENtree as an ontology server:
- Seamlessly access more than 80 public ontologies as well as more than 50 SciBite VOCabs.
- Single API to search, browse, consume all hosted ontologies, whether being a public or a corporate one.
- Unique authentication mechanism whether being the default one or the one already used within your organization!
- No need to monitor underlying public websites for possible new versions. In CENtree, public ontologies are automatically updated every 2 months which also means no download required.
- No ETL (Extract, Transform and Load) process prior ontologies utilization, again, all those possible preliminary steps are done by the SciBite Ontology Team. Indeed, if the ontology or terminology is not in a standard format (which is the case for a couple of them including HGNC, MeSH, CDISC SEND, SNOMED CT, MedDRA, QUDT, Cellosaurus), then, it is transformed into a format loadable in CENtree, usually OWL.
- Validation checks are performed by the SciBite ontology Team. If issues are detected, they are reported to the source ontology provider. In such cases, we do not guarantee a timeline to repair.
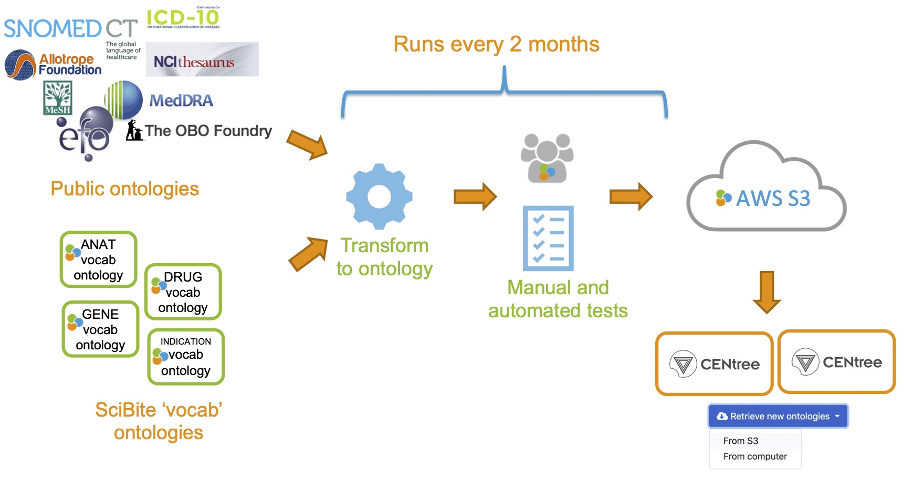
Figure 5: Process to publish public ontologies and SciBite VOCabs in the CENtree UI.
Related articles
In addition to the above benefits, please read our related blog post “SciBite launches SaaS Versions of its Semantic Technology Products.
If you have any questions regarding how CENtree can be used as an ontologies server at your organization or regarding a particular use case, please don’t hesitate to get in touch and one of our experts will get back to you.
Alternatively, download the CENtree datasheet or watch our webinar on Mastering Enterprise Level Ontologies for People and Applications.
About SciBite
Our data-first, award-winning semantic analytics software is for those who want to innovate and get more from their data. Built by scientists for scientists, we believe data fuels discovery and continue to push boundaries with our cutting-edge technology applications and people-first solutions that unlock the complexities of scientific content.
Related articles
-
SKOS in CENtree: Further support in our latest 2.1 release


At SciBite terminologies underpin all that we do. There are many ways to represent and build a standardised terminology, each with different levels of complexity. On one hand you have simple, informal, lightweight terminologies (e.g., glossaries, dictionaries, and thesauri), where the meaning (semantics) of terms is captured using natural language.
Read -
SciBite named ‘Best of Show’ at Bio-IT World 2022 for CENtree


SciBite’s ontology management platform CENtree won ‘Best of Show” at this year’s Bio-IT World 2022. Build the foundations for data sharing with our collaborative ontology management platform.
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.

