A hacker’s guide to understanding bio-ontology jargon
Perfect for those new to bio-ontologies or who work with ontologists - a whole new vocabulary deciphered!

Ironic, isn’t it, that the language used to describe ontologies is often rather loose and differs from community to community? I’m in no way going to fix that problem here. Instead, I thought I’d try and add a bit of light to the shade, (or fuel on the fire depending upon your point of view), and try and pull apart some of the more commonly used bits of jargon you’ll hear people use. This is really my perspective and the way I hear most people use these words.
So if you’re new to bio-ontologies or often hear conversations about them and are unfamiliar with the language, then read on. It’s by no way definitive but think of it as a hackers guide to understanding bio-ontology jargon. I’m also reminded of the longer read available at the OntoGenesis Knowledge blog.
Bio-ontology jargon, lingo, vernacular, lexicon, gibberish (you get the idea)
Ontology
Common synonyms: ontological model
The main artefact that contains all of the classes, textual descriptions and rules.
Ontology class
Common synonyms: ontology concept, ontology term
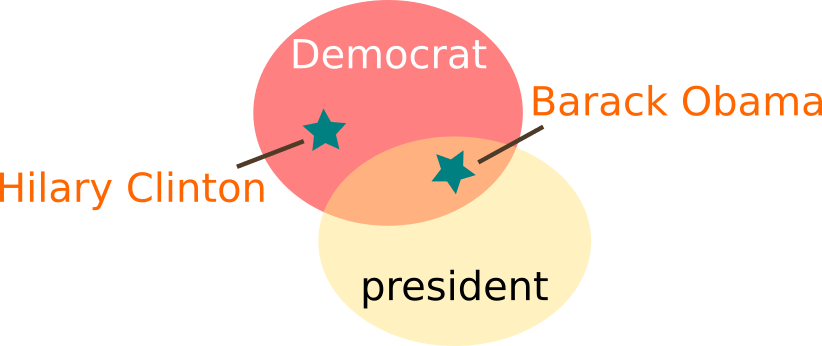
Ontology classes are used to group collections of instances together. These instances are often thought of as the ‘real world’ thing. In the above example ‘Barack Obama’ is an instance of both a Democrat class and the President class, whereas ‘Hilary Clinton’ instance is a member of just the former.
A lot of ontological arguments have taken place over whether to call the primary things that describe entities as classes or concepts, or even more controversially, terms.
Ontology classes and concepts are the bread and butter of an ontology. They are the means by which we describe the entities of interest. An ontology class is a grouping of some set of things by their common properties. For instance, ‘human’ would be such a class. Member of this class would be ‘James Malone’ or, assuming you’re not a bot, you. When you browse an ontology, ontology classes are the things you are seeing arranged in a tree typically. In a very strict sense, ontology term does not refer to the same thing as an ontology class or concept. Term is language used for text in every day use, but you will find most people that talk about ontology terms mean the ontology class or concept.
Primary label
Common synonyms: name, class name, label, class label, preferred label, rdfs:label, skos:label
The name of a particular ontology concept, usually the most common or popular name for the concept in the field. For instance, HIV rather than Human Immunodeficiency Virus. But also Homo sapiens rather than human. Typically, this is dependent on who built the ontology or who the intended users are rather than a single ‘truth’ so don’t be surprised if your favourite label isn’t used. But when people talk about labels they mean the thing it will be primarily called.
Synonym
Common synonyms: alternative term
The alternative words that refer to a class, but which are often not as commonly used. In some cases, this is an arbitrary decision as to which name becomes the primary label and which becomes a synonym. Why? Well, the usage can vary from community to community, although many ontologies mandate a single preferred label so a decision is made. Synonyms should describe the same concept, e.g. Homo sapiens and human.
Definition
Common synonyms: textual definition
The most common use of ‘definition’ is to mean the textual definition of a class, written in human readable language (e.g., English). This is sometimes mixed with the use of the phrase “class description”, but definition is commonly used to mean “human readable definition” in bio-ontologies.
Class description
Common synonyms: class definition, formal class definition, axiomatic description
Confusingly the class description is mostly used to mean the formal set of axioms – the rules – that describe the class in an ontology. For example, the ‘subclass’ relationships a class has to its parent classes or the ‘part of’ relationship such as a digit to a hand. As noted previously, this is different from the human readable definition written in plain language (e.g. English, Japanese, etc,).
Xref
Common synonyms: dbXref, database cross reference, mapping, ontology mappings, class mapping, term mapping, database mapping
Xrefs are mostly used to describe a reference to ‘something’ that is not usually defined in the ontology you are looking at. The original use in OBO (see further down) was for database cross references (e.g. a database record ID), but it is used more widely now such as to reference another ontology (e.g. the ID of a class in another ontology which is a mapping to the one you’re looking at), or a publication reference (e.g. a PubMed ID which may be the source of information for that class).
Most commonly, Xrefs are usually used to map the same concept in another resource, whether it be ontology or database record, for instance BRCA1 ontology class to BRCA1 the gene in gene database. This is useful for tasks involving mapping data sets that are talking about the same thing (e.g. cancer) but have used different resources and therefore different IDs. Xrefs can help automate the mapping of these resources, and hence data.
The use of mapping the same concept is common but not observed universally and is often used to map ‘relatedness’ or even of a relevant subject (such as a publication ID). In the OBO file format (see below) there are explicit mechanisms that enable Xrefs to be treated as ‘equivalent’, i.e. that an Xref is definitely the same concept as the class it is describing. The lesson here is to be a little careful with how you use Xrefs.
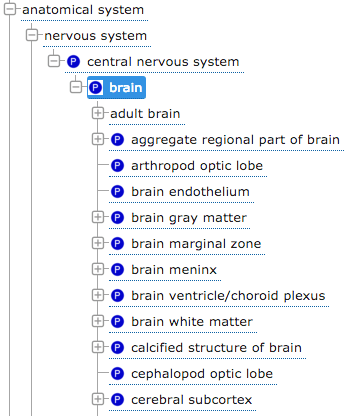
The above tree illustrates two different types of ontology relationships.
Here you can see the subclass relationship (the default indentation in the tree) and the ‘part of’ relationship, shown by the blue P symbol.
We can see ‘brain’ is part of the ‘central nervous system’, and that ‘adult brain’ is a subclass of ‘brain’. (Image from EBI’s OLS).
Relationship
Common synonyms: Class relationship, class relation, object property
The things described in ontologies (species, diseases, anatomy, etc) are connected in ontologies using relationships. This allows us to make arbitrary statements about the things we describe and build up the model of knowledge that ontologies are intended to describe. For instance, that we need to relate blood cells to the circulatory system, that we need to connect Crohn’s disease to inflammatory disease, that we need to connect the UK to Europe.
We call these connections many things in the bio-ontology world but most common is relationships, followed by more explicit types such as ‘object properties’ (primarily an OWL phrase – see later). A useful ‘good to know’ here is that there is an ontology commonly used in the bio-ontology world called the Relations Ontology which is designed to harmonise usage across a lot of the bio-ontologies.
Class axiom
Common synonyms: Class restriction, existential restriction, universal restriction.
Relationships are consumed in class axioms – the part of a description of a class which formally defines the rules of membership of that class (e.g. that a tumour must have a physical site, or that a company must have a legal address). Crucially, class axioms are written in a form that a machine can interpret them, though they are rendered in language to enable a human to write them, much like writing software code. So humans still have a role – for now… If you are interested in learning more on class axioms, the Protege tutorial is a great place to start.
Subclass/superclass
Common synonyms: Parent/child, subtype, is a, inheritance, up tree/down tree, up hierarchy/down hierarchy, subclass/superclass relationship
The most common view of ontologies is as a tree structured to show the hierarchical relationships of subclasses and superclasses. This conforms to much of the traditional understanding of inheritance, namely that classes that are subclasses of another class inherit the properties of their parent. For example, that ‘prostate cancer’ as a subclass of ‘cancer’, shares the properties of the class ‘cancer’.
There is a lot of language used to describe essentially the same thing which you can substitute for subclass. For instance, prostate cancer ‘is a’ cancer; prostate cancer ‘is a child class of’ cancer. Note, ‘subclass’ is a type of relationship (see above) and the statements involving subclasses are types of class axiom (see above) – it is machine readable and is used to define the membership rules of that class.
Class ID
Common synonyms: Class URI, Class IRI, OBO ID
The unique identifier which identifies a class. In ontologies generally this identifier should be globally unique which is why URIs (Uniform Resource Identifier) or IRIs (Internationalized Resource Identifier) (an extension of URIs which enable more characters to be used such as Chinese characters) are used as they include a portion for identifying the resource on the web. Therefore it is possible to embed a domain into that identifier which the ontology developer owns. Not all IDs are of this form though, OBO IDs being one example. They often come in a shorter form such as GO:0006349 and you will see examples of such IDs in literature. Examples of class IDs which all mean the same thing include: http://purl.obolibrary.org/obo/GO_0006349, obo:GO_0006349, GO_0006349.
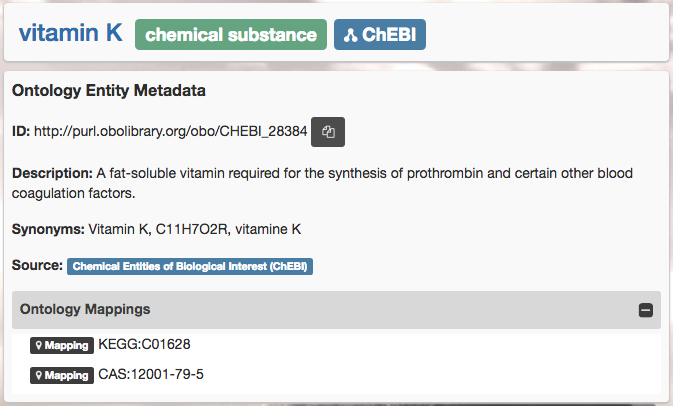
Screenshot from the ChEBI ontology, as shown in Kusp. Annotation properties show human readable text, such as class label (vitamin K), the textual description, and synonyms. Ontology mappings (xrefs) are also shown.
Annotation property
Common synonyms: class annotations, term annotations
The textual parts of an ontology class. This is often human readable parts such as the class label, the class textual definition, synonyms. Note, ontology annotations are annotations which are made on the entire ontology, for instance the date the ontology was created, as opposed to class annotations which are specific to that class (e.g. class label ‘prostate cancer’).
Obsolete class
Common synonyms: obsolete term, deprecated class, deprecated term, obsolete concept
Classes in ontologies, ideally, should never disappear. This is because once they become used, such as for describing a data set, you want the identifier that goes with that class to persist somewhere so there is an audit trail. If the class disappears then it’s harder to know what a person meant when they used it previously. For this reason, classes are instead made obsolete in the bio-ontology world.
They are sometimes removed from the otology and made available separately or, most commonly, included with the ontology and marked as obsolete using some mechanism such as making it a subclass of an ‘obsolete class’ or by tagging it with an annotation. So obsolete classes are classes that were once in use but should no longer be used. Increasingly common is that the obsolete class contains a pointer to another class that should be used instead.
OWL
Common synonyms: OWL file, OWL ontology, Web Ontology Language
OWL (Web Ontology Language [sic]) is a W3C published language for representing ontologies. It contains a specification for defining ontology components such as classes, relationships, and textual properties. It can be represented (serialised) into several formats such as in XML or Turtle (the latter being a more human readable form of the language).
So an ontology can be created in OWL, the language, but saved into formats such as XML and Turtle. Confusingly, it is common to hear of reference to ‘an OWL file’ even when the file is a Turtle or XML file; here they mean the underlying language rather than the format it is stored in on file.
OBO
Common synonyms: OBO file, OBO format
OBO – originally Open Biomedical Ontologies, but which has evolved to Open Biomedical and Biological Ontologies – is commonly talked about in two contexts, both related. The first is the OBO file format. Like OWL, OBO is essentially a language used to describe an ontology, the primary difference being OBO was intended for bio rather than a broader all encompassing language, such as OWL. As such, it began life as a simpler language than OWL but has grown in scope over the years as ontologies became increasingly complex and has evolved to become an official subset of OWL.
The second context is in the OBO Foundry, a collection of communities that are attempting to coordinate how they develop their ontologies, such as using a common framework, common relationships names, common annotation properties, and so on. Obviously the OBOs here are not one and the same, though there is a strong relationship (most of the same people are involved in both). As a rule of thumb, most of the time people saying OBO mean the file format, and will use the phrase ‘OBO Foundry’ (or sometimes just ‘the Foundry’) to refer to the collection of people.
BFO
Common synonyms: ‘Buffo’, Basic Formal Ontology, the philosophy bit
BFO (Basic Formal Ontology) is an upper level ontology used by a growing number of bio-ontologies. It has grounds in realist philosophy (which I absolutely will not go into here) and contains the very high level concepts used to group the more specific things you may be familiar with (such as people and instruments). For instance, BFO contains a class ‘occurrent’ which is, very loosely, stuff that happens over time.
Under ‘occurrent’ would be classes such as ‘behaviour’ and ‘cell growth’. BFO is used to help provide a common structure to the ontologies so they can all interoperate (i.e. all the ontologies use the same class for grouping ‘things that occur over time’). If you are not interested in developing ontologies this is probably as much as you need to know about BFO.
ICBO
The International Conference for Biomedical Ontologies. The main international conference focused on bio-ontologies. There is also a longer running side meeting at ISMB which predates ICBO called Bio-ontologies which is also well attended.
BioPortal and OLS
BioPortal is an ontology repository and browser hosted at Stanford University. OLS (Ontology Lookup Service) is an ontology repository and browser hosted at the European Bioinformatics Institute. Both have similar aims in allowing users to query, browse and, through APIs, programmatically access bio-ontologies. The OLS has a more restrictive set of ontologies loaded into it – specifically those used at EBI and by their service teams, whereas BioPortal has a larger, wider remit whereby any bio-ontology can be uploaded and served from the service, including self-submitted ontologies.
Related non-bio-ontology jargon
There are also a bunch of other terms you’ll hear in the community which are not specifically about ontologies, but are relevant. This list is large so I won’t cover it all here, but they include:
Vocabulary
Common synonyms: terminology
Vocabularies are bags of words that are put together to encapsulate the language used in a domain. Indeed, much of the tooling we develop relies on this very process – consuming semantics of an ontology and turning them into vocabs for the purposes of entity extraction and tools for editing those vocabs.
SKOS
SKOS (Simple Knowledge Organisation System) is a specification that can be used to define a basic structure of things in a domain. The strong semantics you find in an ontology is weakened significantly in a SKOS model and this is a key differentiator between the two. SKOS provides a model for going up and down a tree in a hierarchical manner through a ‘broader than’ and ‘narrower than’ relationship pairing, however there is no implication of inheritance here, only directionality. It is common that SKOS in bio-ontologies is used in this manner, but it is not a requirement.
So there you have it, a rough guide to bio-ontology jargon. Hopefully it clears the muddy waters a little when faced with an onslaught of unfamiliar terminology.
To find out more about how the SciBite Semantic Platform can transform your data, or get in touch with the team.
Related articles
-
Are ontologies relevant in a machine learning-centric world?
SciBite CSO and Founder Lee Harland shares his views on why ontologies are relevant in a machine learning-centric world and are essential to help "clean up" scientific data in the Life Sciences industry.
Read -
How ontology enrichment is essential in maintaining clean data
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the final blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.