SciBite & RDF (Resource Description Framework) – A natural semantic fit
In this article, we’ll explore how SciBite’s platform works with the semantic web and its primary data representation format, RDF, along with the benefits each technology brings.

Anyone who reads our regular blogs will have seen the word “semantic” is pretty important to us! We often stress how the word itself does not imply any particular data format but describes how we focus on extracting meaning from textual data. For instance, rather than just storing the characters a-s-t-h-m-a, with SciBite’s platform, a computer can understand this as an instance of the concept of asthma, an inflammatory disease of the lung, with all of the subsequent value that brings.
In turn, this leads to more powerful search, discovery and analytical capabilities. However, the word “semantic” can be a bit of a loaded term and for many in the technology space, it has a strong association with the semantic web, and its primary data representation format, RDF. In this article, we’ll explore how SciBite’s platform works in such an environment and the benefits each technology brings.
Utilising resource description framework primer
RDF (or Resource Description Framework) is simply a way to represent data, just like spreadsheets, databases, PDF, XML and many other formats.
Unlike these formats however, RDF is specifically designed to store meaning alongside the data. Instead of an element labelled “Bob” (meaningless to a computer), a “tag” (more formally known as a URI) that identifies this as a Person object, according to some agreed reference (known as an ontology). Presented with 1000s of different RDF documents containing Person tags, the computer can identify all people and perform calculations like population analysis without any manual input. RDF data is written in the form of “triples”, where everything is represented in a subject-predicate-object type relationship (for example, SciBite subject -> is_based_inpredicate -> Cambridgeobject). When combined with ontologies, RDF has huge potential for powering integrated data analytics and has many applications within pharmaceuticals, healthcare and life sciences.
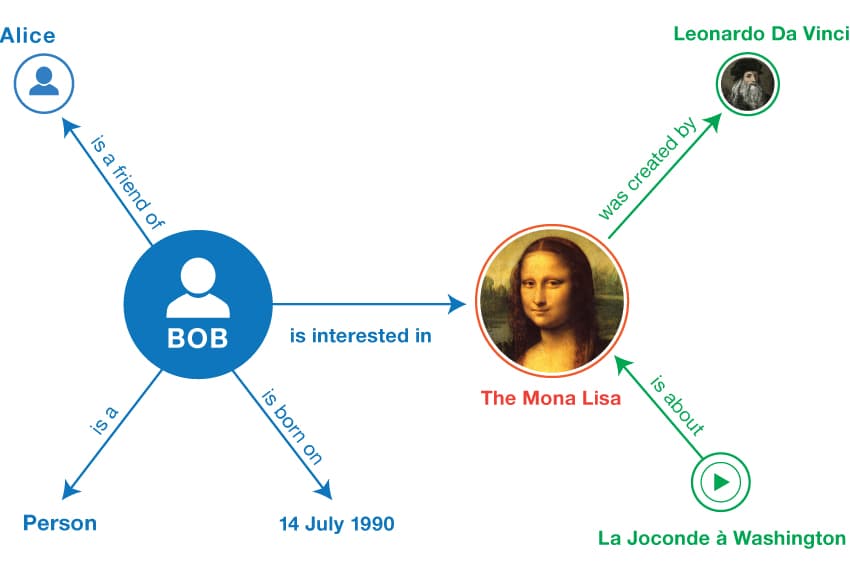
The figure depicts an example from the W3C website, describing a person “Bob” and his interest in a famous painting. Each fact is represented by a subject and an object, being connected by the arrow representing the predicate. Thus, the entire graph can be seen as a set of such triples.
Nanopublications: The extra layer for your data
One of the most critical concerns of those of us who work with pharmaceutical data is that of provenance. Where exactly did the data come from, who made it, what instruments were used etc. There is overwhelming evidence of how millions of dollars have been wasted in drug-discovery efforts due to poor data provenance. This issue is fundamental to all areas of science and technology and led to the development of a proposed solution known as Nanopublication. While not specifically tied to RDF, most nanopublication work to date has been built on an RDF approach as first outlined by Groth et al in 2010.
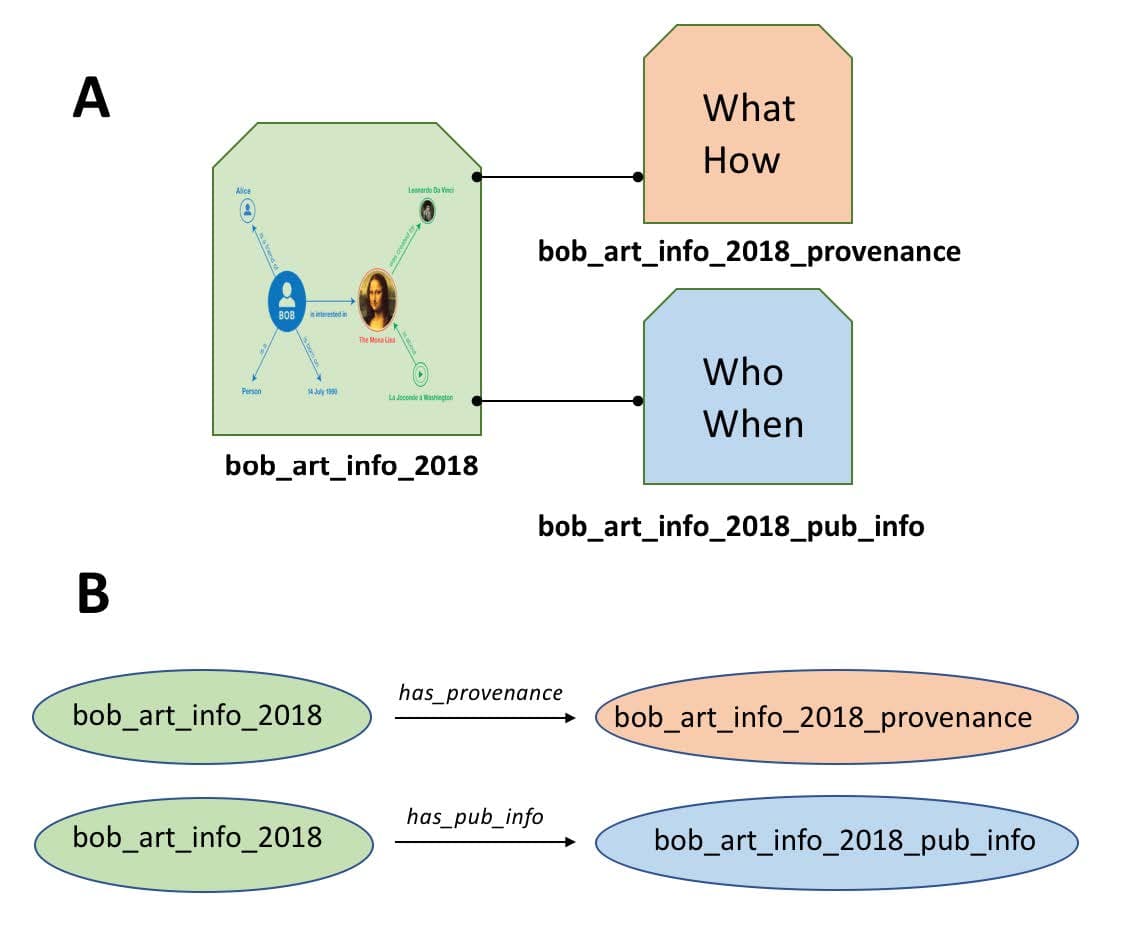
To illustrate how a nanopublication works, we can take the RDF from the Bob/Mona-Lisa example above and put that into a “bucket” as shown in the figure. We give that bucket a unique name such as bob_art_info_2018. Now that the “bucket” containing all the RDF we care about has a name (identifier), we can attach information to the bucket itself.
In the core nanopublication guidelines, we create two additional sets of RDF, one to describe the provenance of the data and another to describe the publication information (where did the RDF come from?). We describe each of these in RDF and put this into new buckets, again, giving those buckets unique identifiers illustrated in part (A) of the figure. Now, we just need to generate a couple of more very simple RDF triples to connect our provenance and publication info buckets to our original data (B). Now there is an unbreakable assertion between the data and its provenance, which is the foundation of good data stewardship so desperately needed in today’s data-driven environment. Technically, nanopublications are achieved by representing the data in quads, rather than triples, with the fourth data element representing the identifier of this “bucket”, more formally known as the named graph.
The role of SciBite’s Semantic Technology
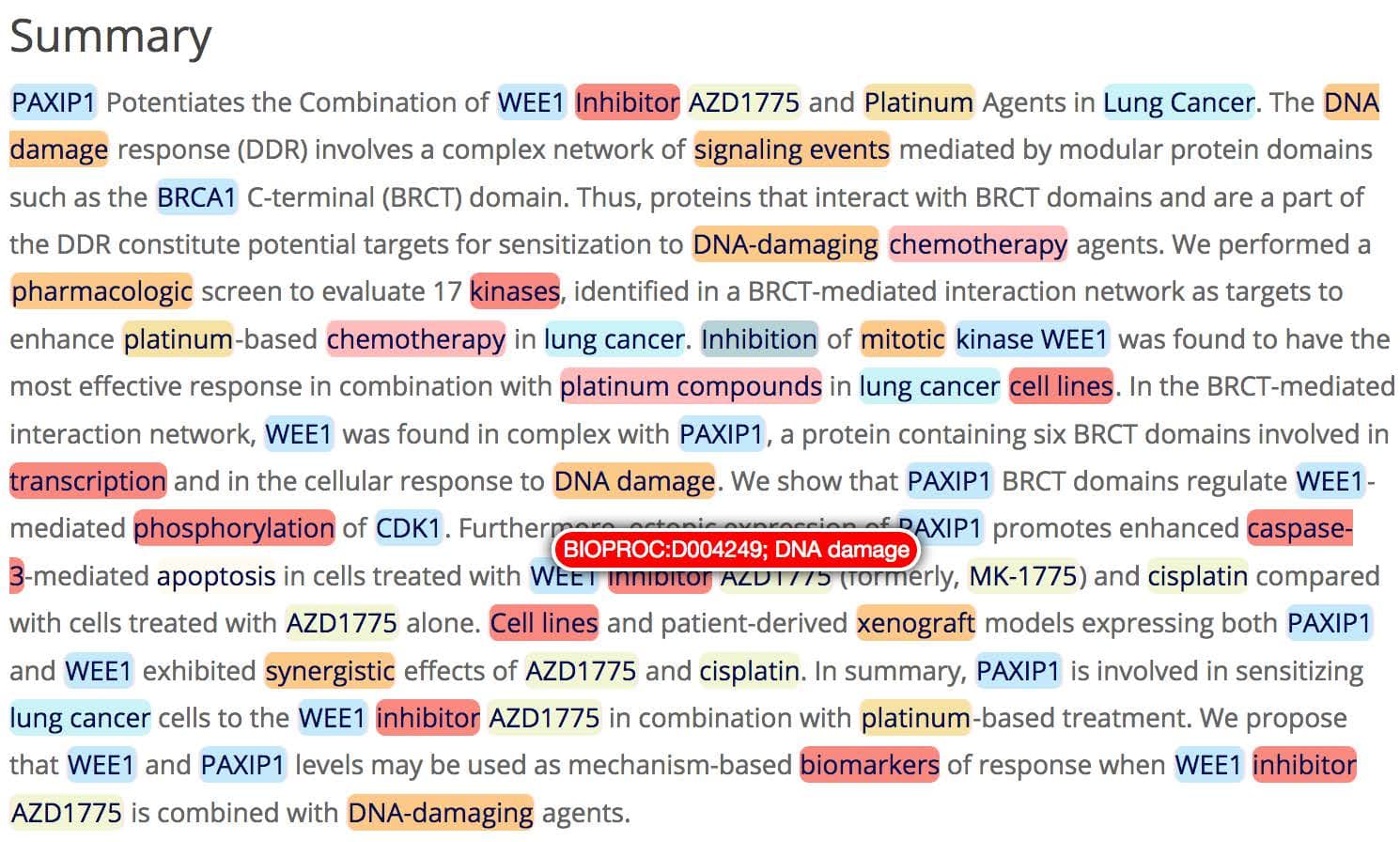
The primary role of RDF is to represent structured data in a way that is meaningful to computers, enabling subsequent discovery and analytics. An obvious question is how RDF can be utilised for unstructured textual data, such as patents, publications, internal documents and so on. It is here that SciBite’s semantic platform provides the necessary bridge between these two worlds. With our platform in place, customers are able to convert possibly millions of text documents into structure data, including RDF, for inclusion into their data environments. This brings enormous analytical power, for instance, combining internal biological assay data with millions of facts extracted from the public literature. The image below shows a typical analysis with TERMite, our award winning named entity recognition engine. TERMite has identified many important concepts, enabling the generation of rich relationship data from an otherwise poorly represented text document.

The SciBite platform achieves this through a number of unique features available in a system that was specifically designed to provide the ‘semantic glue’ to join disparate data. The figure below outlines the different modules of relevance in our highly customisable platform, and why they matter for this use case.

- Connector: SciBite’s Connector Framework (SCF) allows for the routine, automated extraction of new data from specific sources both internally and on the internet
- Parser: SciBite has over 120 parsers for different data sources, including many of the key public and commercial data sources. Where a parser does not exist, they can be rapidly prototyped using our own simple parsing language, Java or Python
- NER: Named Entity Recognition. SciBite ships with over 100 of the most important ontologies for life-sciences/pharma. This allows the system to generate high quality relationship graphs between genes, diseases, drugs etc within or across document sources
- NLP: Natural Language Processing. Beyond entity recognition, customers can devise specific linguistic searches to look for events such as drug-side effects, protein-protein interactions, biomarkers and many more
- Workflow: Users can insert custom-processing code within the server that allows analytics to be performed in-situ and returned in a standard way via the SciBite APIs. We use these “workflow” plug-ins to tailor the pipeline for specific use-cases, as well as calculating things like relevancy scores, which identify the most important entities mentioned in a document
- Writer: It’s all very well being able to generate data, but often it can be difficult for users to consume this into RDF systems without extensive post-processing. The SciBite platform is different, as described below, the system supports whatever form of RDF you wish to create
Flexibility for your unstructured data
While RDF is an important step forward in data standards, it does not force developers to use any particular schema to encode the data. For instance, let’s say you wish to represent the phone number for a particular person. Perhaps the simplest and most obvious way to represent this is shown in (A) in the figure, a single RDF triple which does the job nicely.

However, we might want to represent multiple numbers for Bob. The RDF could be changed to the form shown in (B), allowing us to represent this perfectly well. Alternatively, we may need to represent the data in a more sophisticated manner, such as that shown in (C). While this latter example might seem to be over-complicating things, it more accurately represents the way complex data will be encoded in RDF in real world use-cases.
The question here is not which of the 3 models (or ‘schemas’) above is correct, neither is more correct than the other. The point is that the choice of model and predicates (which are sourced from ontologies) is pretty much down to the developer, designed to meet the needs of that use-case. Thus, SciBite’s platform does not emit a specific form of RDF but uses a templating system whereby the developer can specify the skeleton of the RDF, using placeholders for live-data values from each document that is searched. This is critical in modern-RDF processing.
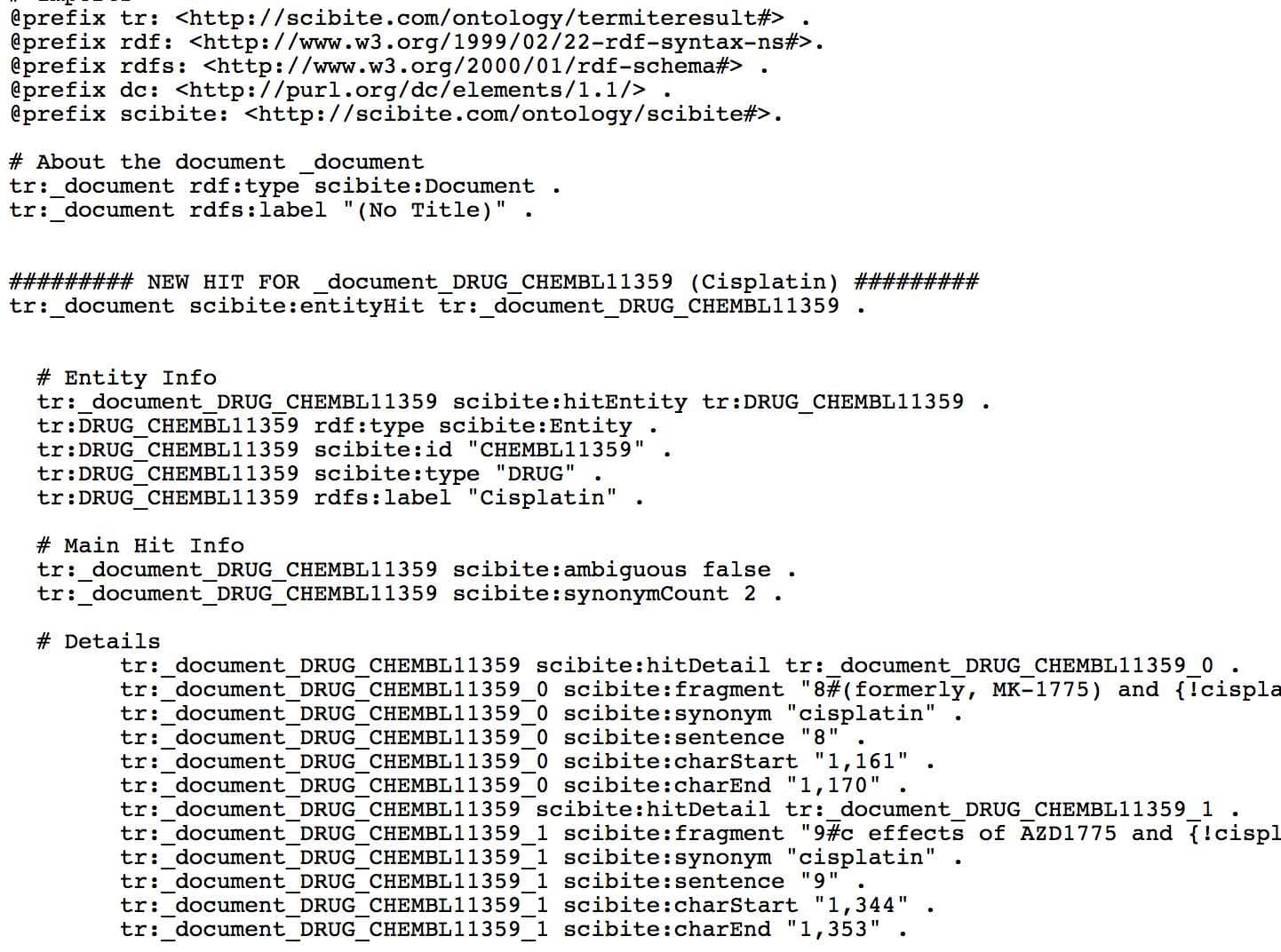
The figure shows the default RDF emitted directly from the SciBite system, representing which entities from various primary ontologies are found in which documents. This can be easily imported into a tripe store, allowing users to query a massively enriched knowledge graph, ensuring access to important findings in their analysis. And, because the SciBite platform is specifically designed for systems integration, you can create an Extraction-Transformation Load style pipeline to generate powerful RDF assertions with ease.
The output shows basic triples rooted in some basic ontologies for representing the data. However, by editing the template file in a text editor, one could easily change this to generate nanopublications in an n-quads style output, or use different predicates or ontologies as required. An example of a nanopublication template is shown in the figure. It is a simple text-file where placeholders capture important data from SciBite’s analytics. Of particularly utility is the ability to use many different URIs within the system. Thus, if you are marking up diseases and require the output to use SNOMED or ICD identifiers for instance, that is a simple setting in the workflow.
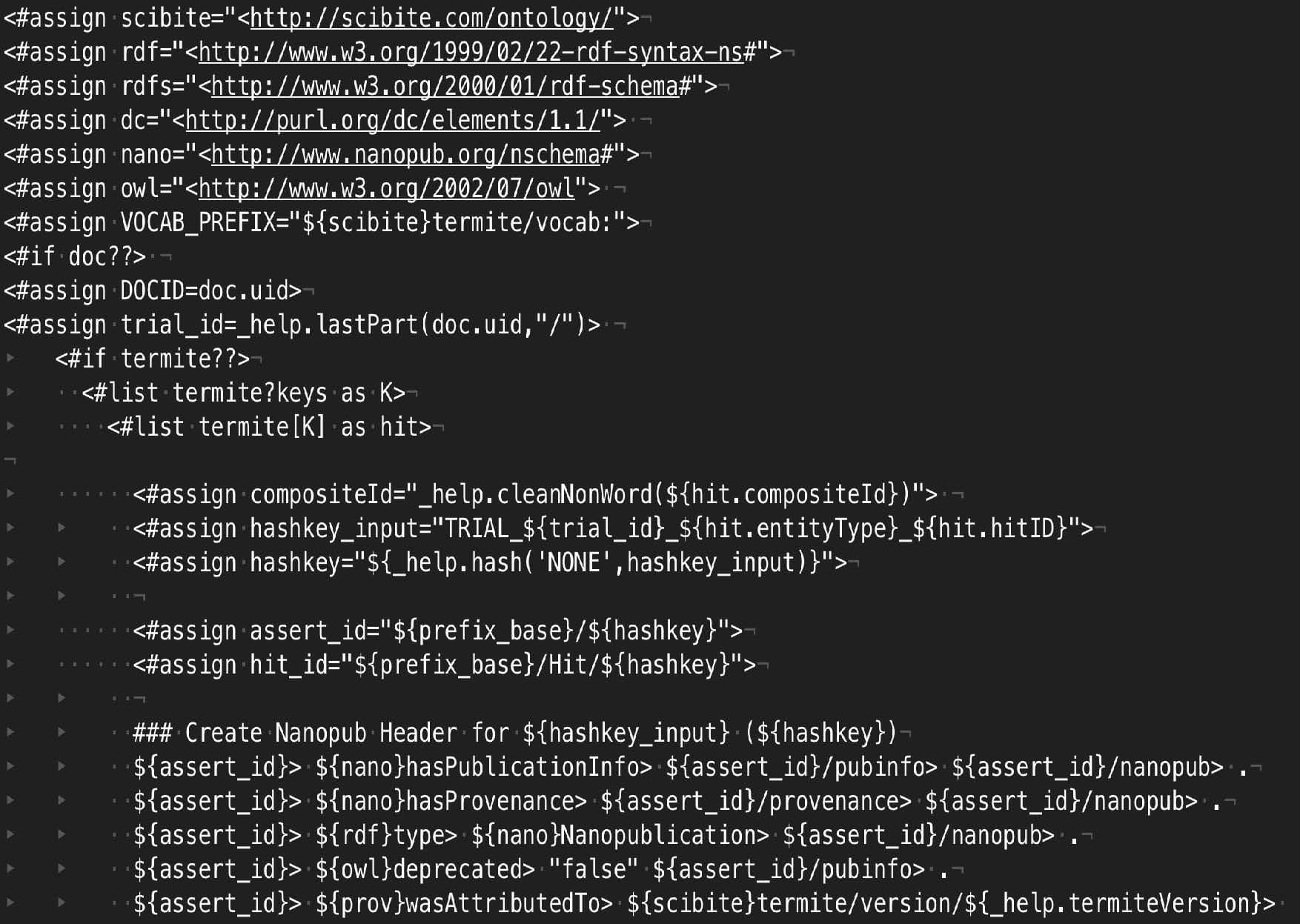
Apart from the obvious abilities to connect to many different data sources and perform high quality NER/NLP, a critical consideration must be the ability to easily customise the RDF to the needs of the target system without the need for costly software engineering projects. The SciBite platform provides all this and much more, bringing a new dimension to your semantic technology projects and delivering some of the values outlined below:
| A Better Search Experience | Find data that mention “Lipitor” when the search term is “Atorvastatin” |
| Concept-Type Searches | Find data that mention a gene or indication and another topic of interest |
| Connections | Find data like “What are all the targets connected to drug repurposing or a particular genetic variant or indication” |
| Fact Extraction | Find all relationships or values e.g. drug-side effect or dosage values or inclusion/exclusion criteria |
| Ontology Queries | Find any data that mention a kinase or inflammatory disorder |
| Transformative Data Integration | Add structure to unstructured textual and connect it to databases and other systems to provide a complete view across the organisation |
Generating RDF from unstructured documents is only one aspect of the capabilities of the SciBite platform. Scalability challenges of processing millions of documents are tackled through a combination of basic speed (the system can process over 1 million words/second) and parallelisation through technologies such as Hadoop and Elastic Search. In addition, SciBite provides machine-learning based solutions to extracting specific facts from text documents, such as drug-adverse events, sentiment analysis or detecting novel technologies. The ability to embed a semantic enrichment platform directly into your data processing pipeline is a must for today’s data-driven science, ensuring vital knowledge is no longer lost.
For more information on how the SciBite platform can work with your RDF project, please contact the team.
Related articles
-
Large language models (LLMs) and search; it’s a FAIR game


Large language models (LLMs) have limitations when applied to search due to their inability to distinguish between fact and fiction, potential privacy concerns, and provenance issues. LLMs can, however, support search when used in conjunction with FAIR data and could even support the democratisation of data, if used correctly…
Read -
A review of the Pistoia Alliance Spring Conference 2023


Last week SciBite was lucky enough to attend, and present at, the Pistoia Alliance Annual Spring Conference ‘23, held at the fantastic Leonardo Royal Hotel, St. Pauls, London. Read the thoughts of our Director of Technical Consultants, Joe Mullen
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.

