Using ontologies to unlock the full potential of your scientific data – Part 1
In the first of this two-part blog, I describe what ontologies are and how you can use them to make the best use of scientific data within your organisation.

The data challenge
The struggle to effectively utilize the increasing volumes of data available is a common challenge in the Life Sciences research industry. Over the last 20 years, new technologies are revolutionizing biological research and its applications. Whereas once we had only sequence data, we now have expression data, metabolomics, metagenomics, and variant data, with whole-genome sequencing and single-cell sequencing just on the horizon. With that huge growth in data comes the need to be able to search, integrate and analyse it in scalable ways.
Because it is now easier and cheaper to generate and store ever-greater volumes and types of data, the biggest barrier to exploiting it effectively is now the quality of its metadata. Computational models, such as Machine Learning and Artificial Intelligence, can only be as good as the data that has gone into them.
I’m sure you are probably familiar with at least one of the following data issues:
- Large amounts of unstructured legacy data, such as text documents or experimental data with either no or poor metadata
- Data is stored across the organization in silos, with similar information managed in a disconnected, unsynchronised, e.g. separate Species lists in your ELN, LIMS, and other databases
- One or more internal, home-grown vocabularies that have evolved over time and are not mapped to a public ontology
- Lack of controlled metadata entry, resulting in what is there being patchy and unstandardized
Each of these issues alone presents a challenge to finding information. Any combination of them and it becomes almost impossible.
To illustrate this problem, the figure below lists a selection of the many varied words and phrases scientists submitters to the EBI’s ENA resource [1] use to refer to the term ‘male’.

A selection of the many varied words and phrases scientists submitters to the EBI’s ENA resource
At SciBite this is something we see a lot. In this blog, I’m going to explain how applying ontologies and standards can address this problem and:
- Enable you to share your data more easily across an organization
- Make your data interoperable by normalizing to ontologies both within an organization but also potentially with external data, such as the Gene Expression Omnibus (GEO)
- Increase your chances of spotting errors and inconsistencies, such as missing and incorrectly labeled data
- Harmonise and clean your data so that it can be read and understood by computers, providing a foundation for technologies such as machine learning and artificial intelligence
- Make your data Findable, Accessible, Interoperable, and Reusable – the key principles of the FAIR data movement [2]. I’ll come back to FAIR again in a moment…
What is an Ontology?
Let’s take a step back for a minute and talk about what an ontology actually is. Ontologies serve a vital role in providing an open, consistent, stable identifier for a given “thing” and developing consensus in the scientific community regarding what that ”thing” is. Ontologies also describe how types, or classes, of thing relate to one another. Essentially, they encapsulate the scientific knowledge in a given scientific domain. That is one of the reasons why building ontologies is so difficult – you have to get scientists to agree with each other!
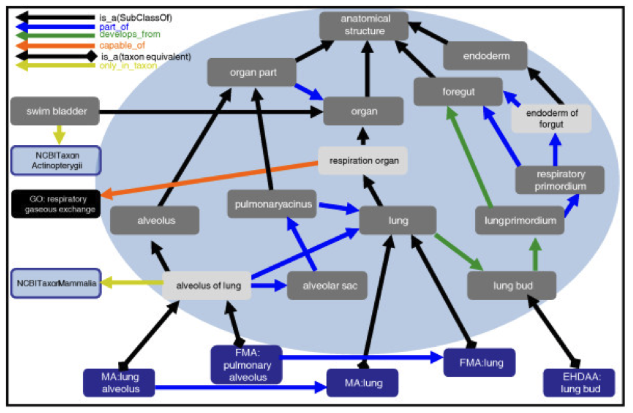
Figure taken from https://genomebiology.biomedcentral.com/articles/10.1186/gb-2012-13-1-r5
Importantly ontologies are created by humans but readable by machines – they provide a way for us to explain to a computer how humans understand the things that exist in the world and how they interrelate.
Ironically and confusingly, the word ‘Ontology’ often gets used ambiguously to mean any kind of controlled vocabulary, list, or taxonomy. In fact, these artifacts might be thought of more usefully as being on a spectrum with increasingly strong semantics, from a collection of terms (tags) to enhance categorization (a ‘folksonomy’), through to a formal description of a domain with classes and relationships.
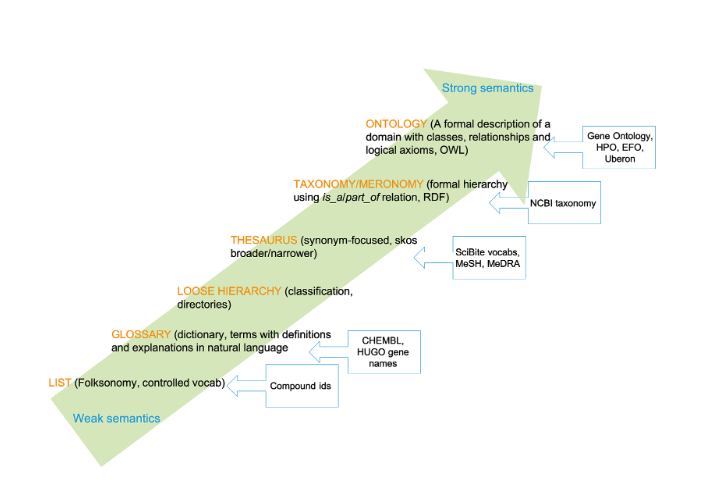
There’s no inherent value judgement about whether one of these is better than another, it’s more a case of choosing the right tool for the right job. For example, there’s no reason why your shopping list needs to be fully axiomatized or even have a hierarchy.
Why use standard ontologies?
So you can see that by using an ontology term in your system, you are doing more than just using a standard name and ID. Adhering to a standard is massively useful in itself because it enables you to align your data both inside and outside your organization. That’s the Interoperable part of FAIR. But beyond this, an ontology class actually means something.
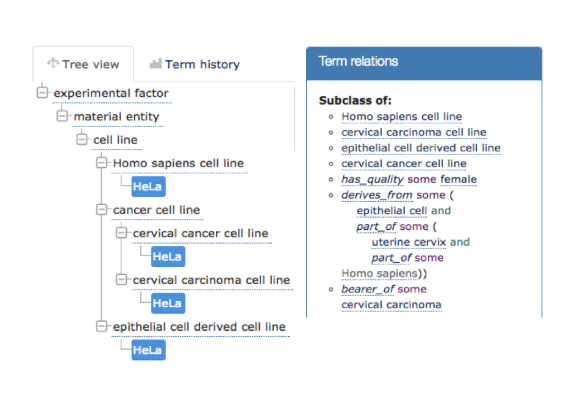
For example, by annotating to HeLa cell line from EFO, you get all of this other information for free such as that HeLa is a cervical cancer cell line, derived from epithelial cells. By using an ontology, you have access to all of this additional knowledge that has been baked in for you by experts, and that’s incredibly valuable. It means you can ask meaningful questions at different levels of granularity by leveraging the hierarchy and query based on relationships such as “Which targets are associated with inflammatory disorders?” and “Which other targets in the same class as a target of interest and what diseases are associated with them?”
Ontologies are the key to unlocking FAIR
Anyone producing or managing data should be acutely aware of whether it will be Findable, Accessible, Interoperable, and Reusable (FAIR). For example, a recent report has identified clear financial penalties as a consequence of not having FAIR research data [3].
FAIR is about making data reusable, but how can you do this if we don’t know if we’re talking about the same thing? For example, an investigator searching for articles relating to ‘myocardial infarction’ would miss references to ‘heart attack’ or ‘serious heart event’. Thus, ontologies are the key to unlocking FAIR – data can only be reused if it is well-described and of high quality.
While FAIR started as FAIR is by no means limited to academic research. For example, we’ve worked with Bristol-Myers Squibb to bring smart data entry to assay registration. Watch our webinar on building a common scientific understanding with FAIR data.
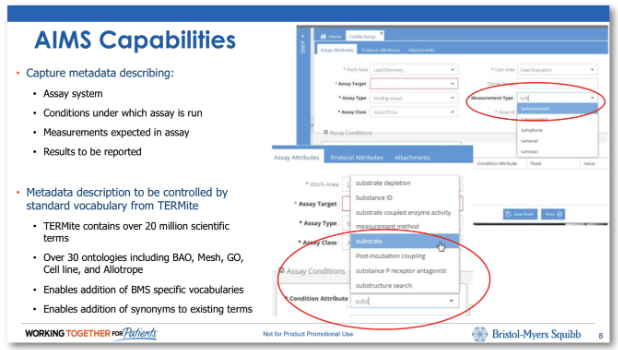
This application uses ontology-driven data entry through standards such as the Bioassay Ontology (BAO). This ensures that users enter high quality experimental meta-data from the outset and puts the user experience front and centre by avoiding horrible ontology tree navigation panels, and using smart, helpful autocomplete systems instead.
Ontologies aid Machine Learning and AI
Ontologies can be really valuable in strengthening Machine Learning (ML) and Artificial Intelligence (AI) strategies. Without using ontologies, an AI model needs to learn everything from scratch. Think of it like going into a library to find a book without some form of classification or indexing to guide you to its likely location. The Dewey Decimal system was developed to help people find the right information faster, and ontologies serve a similar purpose in science.
Since ontologies encapsulate a common model of knowledge associated with a given domain, they can give AI a ‘head start’ by identifying what is known in the context of an existing scientific framework. For example, a machine learning algorithm doesn’t need to be ‘taught’ that Crohn’s is an inflammatory disease if this relationship is already encapsulated in the ontology that is used to enrich the input. Similarly, by annotating content with the MedDRA ontology, the annotated concepts will be known to be adverse events, rather than simply predictions of something that might be. Read our use case on how Ontologies and Machine Learning work together.
Ontologies can also help develop training sets for machine learning, such as using UMLS, or other similar large structured vocabularies of medical terminologies, to provide a ‘skeleton’ upon which an AI algorithm can operate in order to achieve its results [4].

Another example is to use categories, such as anatomical location, in a question answer system such that it constrains the answers patients can give [5].

Ontologies can be used to normalize and clean data going into AI and ML so the machine knows what type of entity is being described in the data. For example, does ‘hedgehog’ refer to a small spiky animal or the gene with the same name? Because ontologies are human models and, therefore a gold standard, they can be used to validate the results coming out of ML and AI systems.
Read my next blog, where I’ll describe how to use and choose the right biomedical ontology, and how to build and manage ontologies. In the meantime, why not watch my recent Introduction to Ontologies Webinar.
[1] Courtesy of N. Silvester, European Nucleotide Archive, EMBL-EBI
[2] Wilkinson, M. D. et al. (2016). The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3:160018 doi: 10.1038/sdata.2016.18
Related articles
-
Using ontologies to unlock the full potential of your scientific data – Part 2
This blog post focuses on mapping, building, and managing ontologies. In my previous blog, I described what ontologies are and how you can use them to make the best use of scientific data within your organization. Here I’ll expand upon this and focus on mapping, building, and managing ontologies.
Read -
How biomedical ontologies are unlocking the full potential of biomedical data
Our latest blog explains how SciBite's Ontologies team takes public biomedical ontologies and tailors them so that they can be used for named entity recognition (NER).
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.

