How SciBite technology can facilitate gene-disease relationship extraction
 |
 |
As genomic sequencing technologies get more advanced, large numbers of gene-disease associations have emerged. A gene with an unclear role within a disease is a source of ambiguity and can lead to misdiagnosis. In this blog, we demonstrate how semantic search technology can facilitate Gene-Disease Relationship Extraction.

The problem
As genomic sequencing technologies get more advanced, large numbers of gene-disease associations have emerged. A gene with an unclear role within a disease is a source of ambiguity and can lead to misdiagnosis. Therefore, it highlights the urgent need for a standardized method to evaluate the evidence implicating a gene in disease and thereby determine the clinical validity of a gene-disease Relationship Extraction (RE) [1].
When we use text mining and the Natural Language Processing (NLP) algorithm to extract gene/disease pairs automatically, it is even harder to decide whether the identified pair is a real regulation or a co-incidence. As a result, we are interested in an automatic pipeline for evaluating the clinical validity of gene-disease pairs.
Gene-disease relationship extraction from unstructured text
SciBite offers a wide range of tools for transforming unstructured text into data for down-stream manipulation and analysis. For this piece of work, we used SciBite Search, which uses ontologies to semantically enrich content to enable powerful searches across multiple literature sources.
SciBite Search has a flexible query language that enables users to narrow their search results based on entities, context, and relationships. It also provides metadata that further can be utilized to enrich results. For instance, because SciBite Search has an inbuilt understanding of types of entity, we can search documents for “anything of type gene/protein” in relation to a specified indication. Furthermore, this can be limited to same-sentence co-ocurrence to gain a quick overview of likely gene-disease associations (See Figure 1).
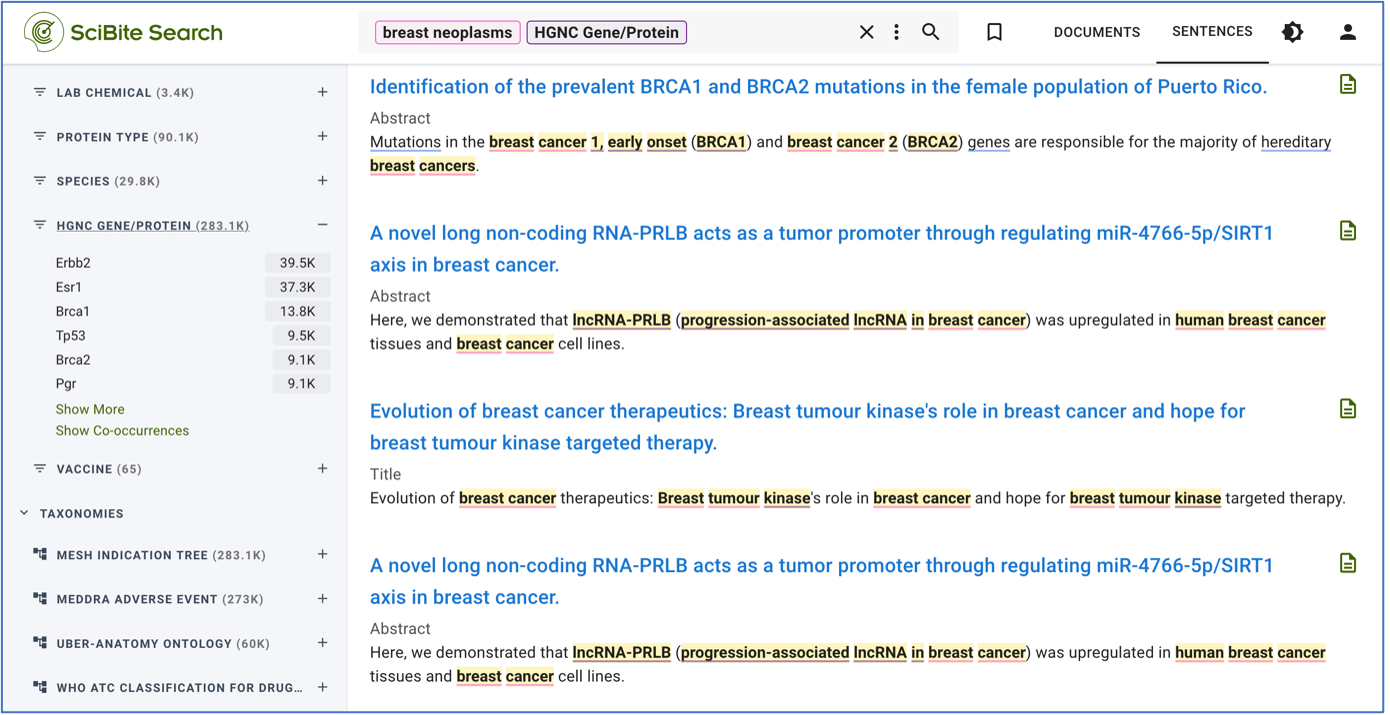
Figure 1. SciBite Search Gene-Disease Relationship Extraction search query. Here we can see sentences relating to breast neoplasms in co-ocurrence with anything defined as a gene or protein.
Now we have a list of sentences that happen to mention some genes and diseases together, as shown in Figure 2. Yet, the question of “are they valid associations” needs to be addressed.

 Figure 2: Sentences with (upper) and without (lower) gene-disease hits. For each sentence pairs of gene/disease (GD) formed the initial list of potential gene-disease associations (GDAs)>(HGNC1101-D001943) and (HGNC1100-D001943).
Figure 2: Sentences with (upper) and without (lower) gene-disease hits. For each sentence pairs of gene/disease (GD) formed the initial list of potential gene-disease associations (GDAs)>(HGNC1101-D001943) and (HGNC1100-D001943).
What is the role of relationship extraction?
For that purpose, we need to perform Relationship Extraction (RE). Relationship extraction is the task of extracting semantic relationships from a text.
In our case, we are interested in importing a sentence with cases of gene-disease and predicting if this cooccurrence is coincidental or describing a real relationship.
Relation extraction can be done in different ways:
- Rule-based models (using language rules)
- Machine-learning approach (training models based on examples)
- Knowledge representation techniques (populating knowledge graphs or semantic nets)
Next step – Knowledge graph population
SciBite offers different technologies to give users various options based on their use cases. The SciBite AI Relationship Extraction model has been trained by our Artificial Intelligence (AI) team, and the model learns to predict gene-disease associations by observing large amounts of data.
On the other hand, knowledge representation approaches (such as Knowledge Graphs) are typically made up of datasets from various sources, which frequently differ in structure and can visualize data dependencies and interactions.
In order to populate a Knowledge Graph, we need to spot our nodes and their relationships. Each relationship can have a weight that indicates its importance. For each GDA candidate, we measure a set of metrics and give each a confidence value (weight). Later we create a knowledge graph using this information in combination with the metadata obtained from SciBite Search.
Figure 3 displays the proposed knowledge-graph schema. The schema of a graph is fundamental; If the schema has been defined correctly, countless inferences can be made based on the data.
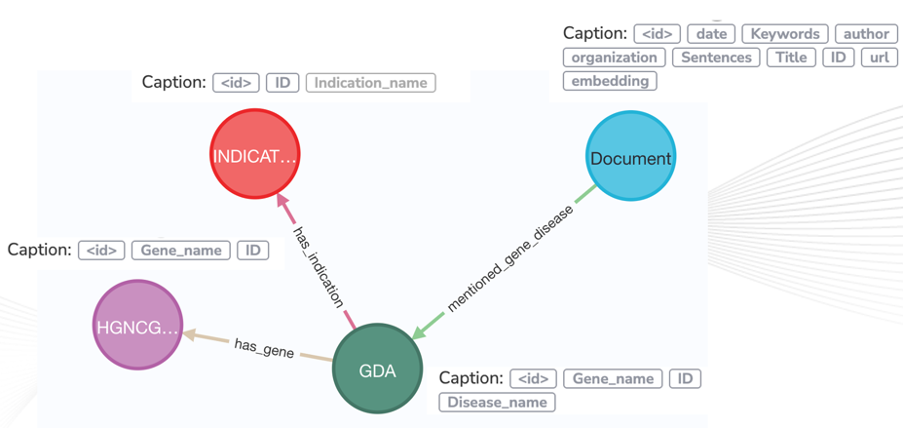
Figure 3: Gene-disease knowledge graph schema. GDA represents Gene disease association nodes, which are linked to gene/protein entities (HGNCGENE), indicaitons and the source document(s) where the relationship was found.
Once the graph has been populated, it is relatively straightforward to ingest additional datasets (via ontology mappings), make changes, and scale the model. Figure 4. Shows a snapshot of the graph.
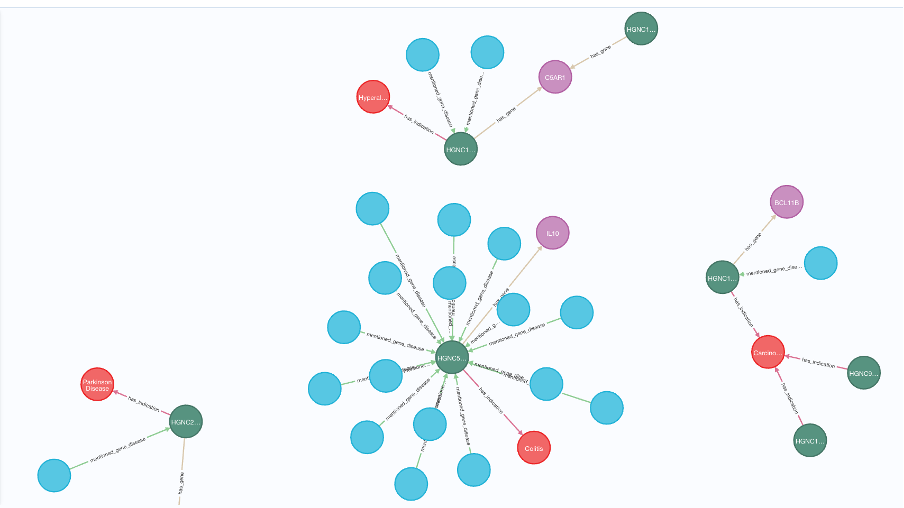
Figure 4: Snapshot of the Gene-disease association graph. (Green nodes: GDAs, Blue nodes: Documents, Red Nodes: Indications, Purple nodes: Genes). This allows the user to explore relationships for any given gene or indication, making it more efficient for a researcher to get an overview of current science for a given topic.
Use cases that a knowledge graph can address
In this section, we cover a few questions that can be answered in SciBite using our knowledge graphs.
For gene/disease association, those GDA nodes that have high incoming cardinality (number of mentions in documents) and high confidences have been proven to be true associations. Those nodes with lower mentions and confidence, and they are neither crowded nor sparce, have been proven to be valid as well and worth investing in as future solutions. Graph data analysis algorithms are extremely helpful in finding this type of node. Figure 5 represents the output of one of the algorithms that we use in SciBite.
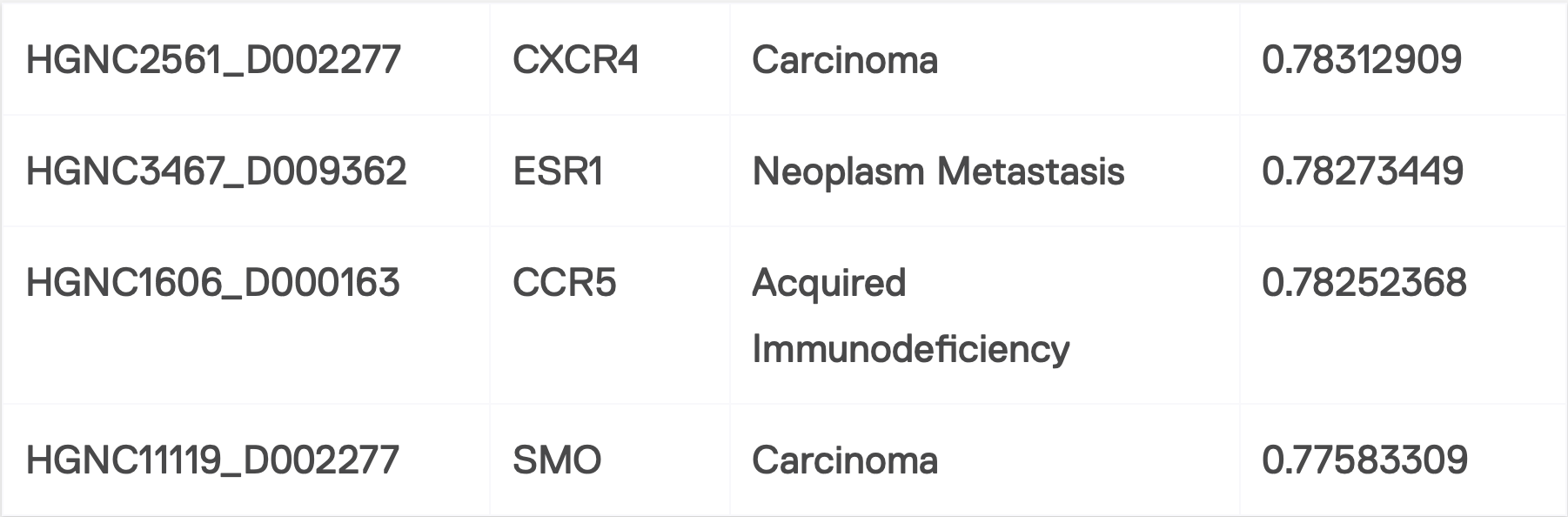
Figure 5: Ranking of GDA nodes base on cardinality and confidence
We also can search the graph for the most mentioned GDAs for a specific disease as Figure 6 depicts.
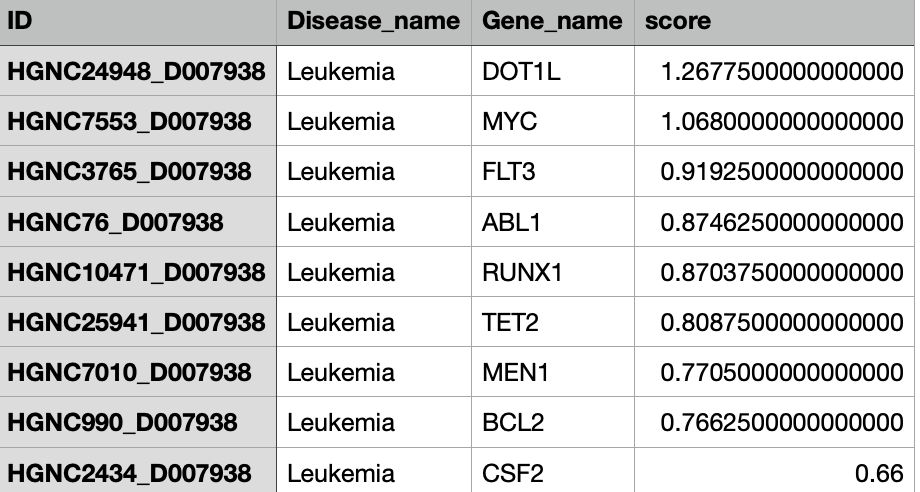
Figure 6a Most Involved Gene In Leukemia
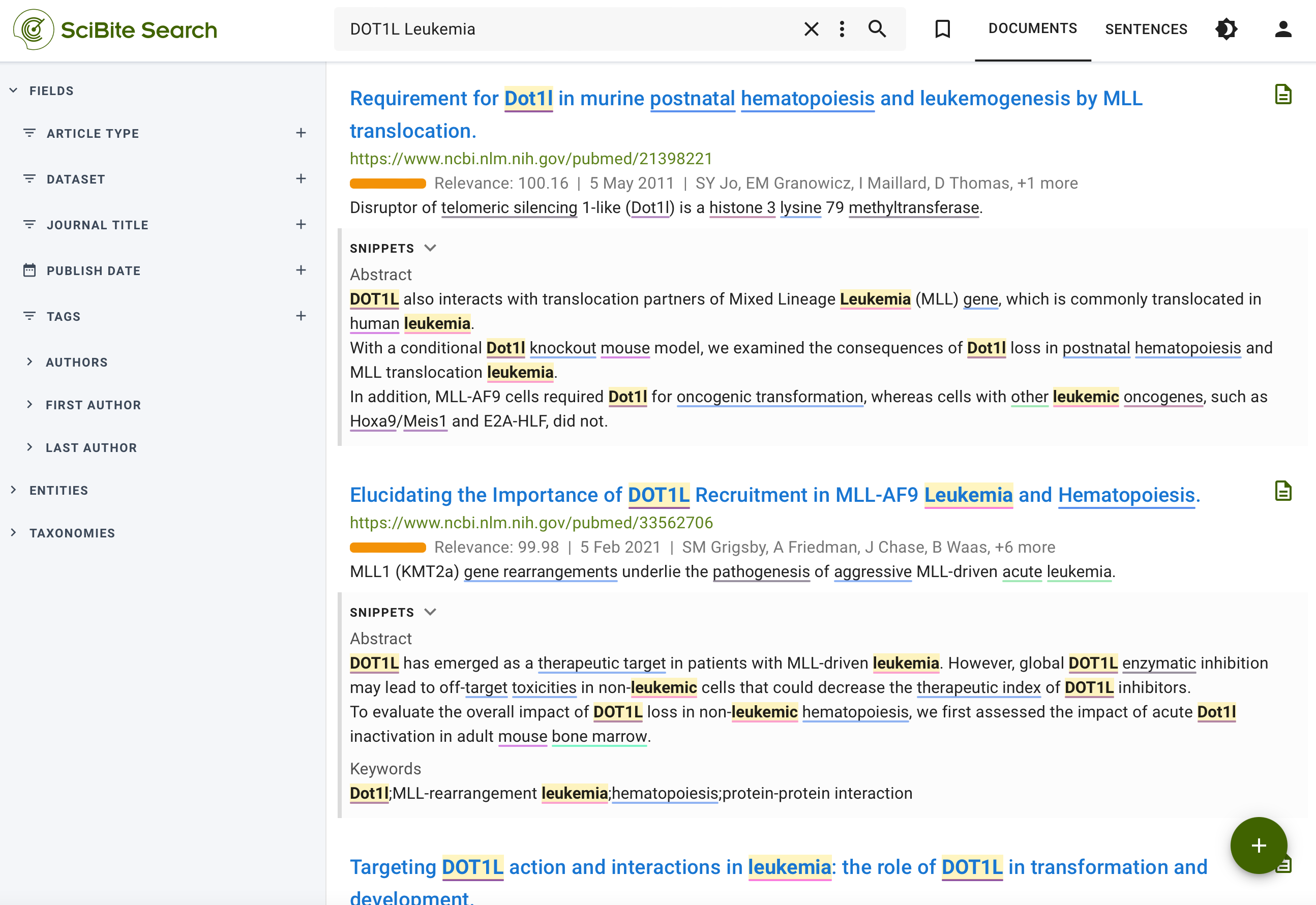
Figure 6b: SciBite Search results.
Finally, we may be interested in figuring out which gene-diseases have the highest mentions in specific organization publications (KOL interests). Figure 7 shows the results of such queries.

Figure 7: Research interests of University of Colorado.
Platform independence
SciBite’s capabilities are completely agnostic to the technology used to represent or store knowledge graphs. SciBite Search, SciBite’s next-generation scientific search and analytics platform offers powerful interrogation and analysis capabilities across both structured and unstructured public data and proprietary sources.
Utilizing SciBite’s robust suite of ontologies, SciBite Search automates semantic enrichment and annotation, transforming unstructured scientific text into clean, contextualized data free from ambiguities like synonyms or cryptic data such as project codes and drug abbreviations. It supports a wealth of features, including marking-up PDF documents, support for federated searches, and natural language queries tailored to the needs of individual users.
Discover more about SciBite Search
References
[1] Strande N. T. et al., Evaluating the Clinical Validity of Gene-Disease Associations: An Evidence-Based Framework Developed by the Clinical Genome Resource. Am J Hum Genet. 2017 Jun 1;100(6):895-906. doi: 10.1016/j.ajhg.2017.04.015. Epub 2017 May 25. PMID: 28552198; PMCID: PMC5473734.
Related articles
-
Addressing common challenges with Knowledge Graphs


In this blog we describe the pivotal role of semantic enrichment in the creation of effective Knowledge Graphs, and illustrate how semantic Knowledge Graphs help answer complex scientific questions.
Read -
SciBite and Stardog build a Knowledge Graph for drug discovery


In a recent webinar, SciBite and Stardog team members demonstrated how to build a knowledge graph to identify candidate drugs for a rare disease.
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.

